Distribution of Rickettsia spp. in Ticks from Northwestern and Southwestern Provinces, Republic of Korea
Article information
Abstract
This study was done to characterize distribution of Rickettsia spp. in ticks in the northwestern and southwestern provinces in the Republic of Korea. A total of 2,814 ticks were collected between May and September 2009. After pooling, 284 tick DNA samples were screened for a gene of Rickettsia-specific 17-kDa protein using nested PCR (nPCR), and produced 88 nPCR positive samples. Of these positives, 75% contained 190-kDa outer membrane protein gene (ompA), 50% 120-kDa outer membrane protein gene (ompB), and 64.7% gene D (sca4). The nPCR products of ompA, ompB, and sca4 genes revealed close relatedness to Rickettsia japonica, R. heilongjiangensis, and R. monacensis. Most Rickettsia species were detected in Haemaphysalis longicornis. This tick was found a dominant vector of rickettsiae in the study regions in the Republic of Korea.
INTRODUCTION
Spotted fever group rickettsiae (SFGR) are obligatory intracellular bacteria commonly found in arthropods such as ticks. Some of the SFGR cause rickettsioses after arthropods transmit them to animals and humans. Common clinical symptoms of SFG rickettsioses are fever, headache, and rash [1]. Currently, SFGR comprise more than 30 species classified into multiple genogroups including: Rickettsia japonica - R. heilongjiangensis; R. massiliae including R. montanensis; R. helvetica including R. tamurae and R. monacensis; and R. akari [2]. Members of the R. japonica - R. heilongjiangensis genogroup have been detected in Japan and the Far East [3]. Specifically, the first clinical case of R. japonica was known in Japan in 1984. It was reported as Japanese spotted fever [3,4]. Since then, it has been detected in Japan, the Philippines, the Republic of Korea, and Thailand [5–8].
R. heilongjiangensis was first isolated from Dysmicoccus sylvarum ticks in Heilongjiang Province of China in 1983. It belongs to R. japonica subgroup of SFGR [10]. Rickettsioses caused by R. heilongjiangensis have appeared in China, Russia, Kazakhstan, and Japan [9–12].
In the Republic of Korea, a variety of SFGR including R. japonica, R. conorii, R. akari, R. australis, and R. monacensis have been reported over 15 years ago [13–18]. R. japonica was first detected from Haemaphysalis spp. ticks in 2003 and human sera in 2004 while R. monacensis was first detected from Haemaphysalis spp. ticks in 2009 [13,14,18]. Additionally, various unidentified Rickettsia spp. were detected in ticks from 5 provinces (including Jeolla-do) during 2011–2013 [19].
Recently, various Rickettsia spp. in other countries have been reported. Nine species or subspecies of tick-borne rickettsiae have been identified in China in the past 30 years [21]. Guo et al. [22] first reported on the existence of R. raoultii in H. erinacei from wild marbled polecat (Vormela peregusna) in China in 2014. It may be assumed that there is a need to examine geographical features (i.g. China-Kazakhstan border) in the identification of various Rickettsia species [21]. Also, since tick-borne disease can be prevalent throughout the country due to climate change, it is important to investigate seasonal occurrence and status of ticks to predict the potential of transovarial transmission [22]. Therefore, the objective of this study was to identify and characterize rickettsiae in ticks collected at different geographical regions in the Republic of Korea.
MATERIALS AND METHODS
Collecting and identifying ticks
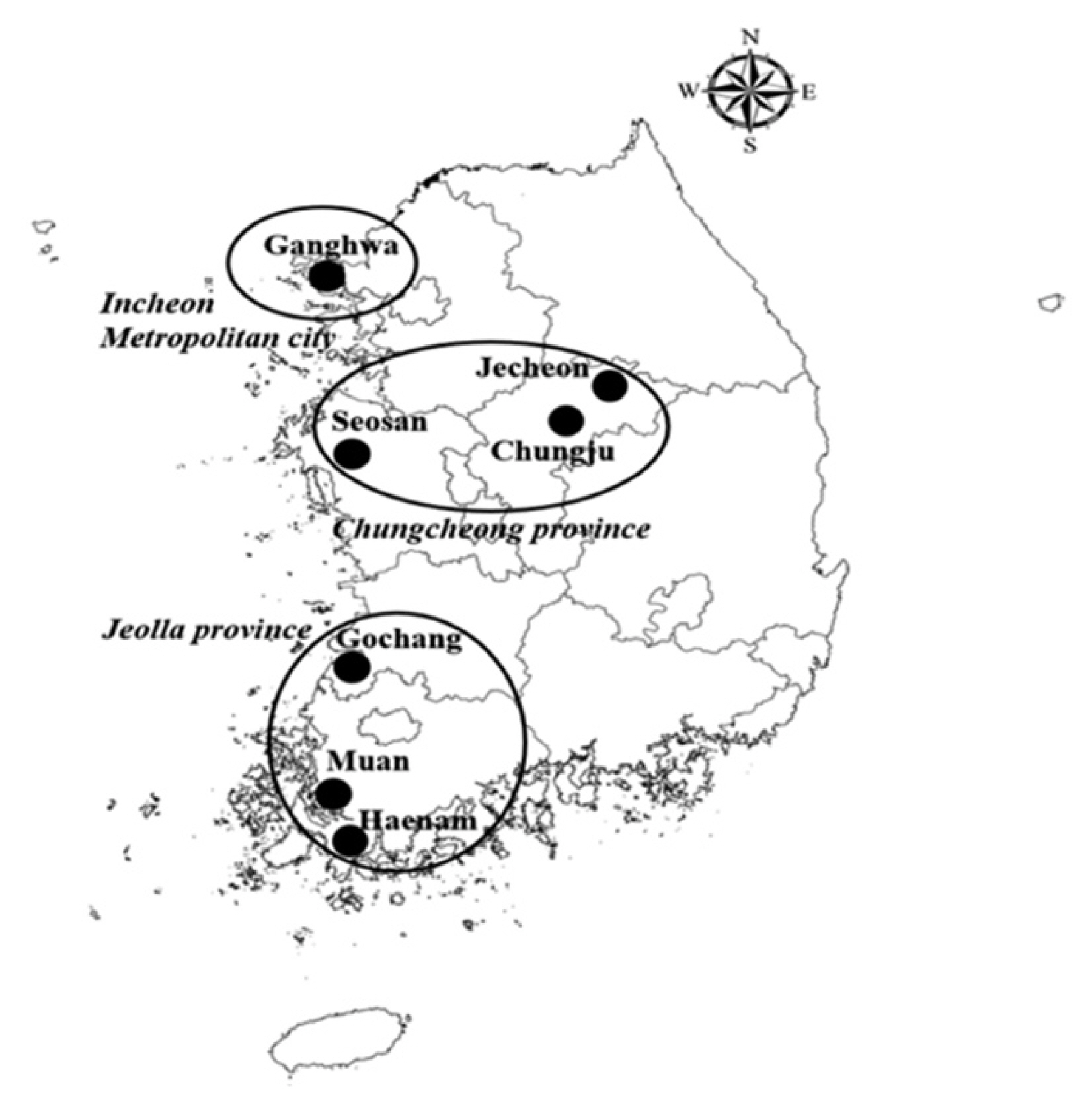
All ticks were collected using tick dragging in the northwestern province (4 regions in Incheon-si, including Gangwha-do (37°44′10.5″N/126°31′47.5″E and 37°45′02.9″N/126°25′26.9″E), Samsung-dong (37°43′47.9″N/126°29′36.6″E), Gilsang-myeon (37°37′31.1″N/126°29′34.1″E), and Bureun-myeon (37°37′04.6″N/126°28′35.3″E)) and 2 southwestern provinces (3 regions in Jeolla-do: Muan (34°51′06.9″N/126°25′03.8″E), Haenam (34°35.68.6″N/126°38.45.3″E and 34°34.01.8″N/ 126°38.16.1″E), and Gochang (36°35′67.6″N/126°33′55.7″E); and 3 regions in Chungcheong-do: Seosan (36°44′26.0″N/ 126°34′05.0″E), Chungju (37°01′43.3′ ′N/127°50′50.0″E), and Jecheon (37°13′39.5″N/128°05′11.5″E) in Republic of Korea from May to September of 2009 (Fig. 1). Ticks were identified and their developmental stages such as larva, nymph, adult male, and adult female were determined under a stereomicroscope. Pooled tick samples were transferred to 2 ml microcentrifuge screw-cap tubes and stored at −70°C.
DNA extraction
Pooled tick samples were washed with 70% ethanol and rinsed with distilled water. Total DNAs were extracted from these samples using G spin total DNA extraction kit (iNtRON, Gyeonggi, Korea) according to the manufacture’s introductions. DNA samples were stored at −20°C until use for DNA amplification.
nPCR to detect rickettsial agents
First, we performed nPCR screening to select positive DNA samples using specific primers for 17-kDa gene: R17K31F (GCTCTTGCAGCTTCTATGTTACA) and Rr2608R (CATTGTCCGTCAGGTTGGCG). The reaction mixture was prepared by adding 2 μl DNA extract and 8 pmole of each primer into a tube of AccuPower® PCR premix (Bioneer Corp., Daejeon, Korea) composed of 1U Taq DNA polymerase, 250 μM dNTP, 50 mM Tris-HCl (pH 8.3), 40 mM KCl, and 1.5 mM MgCl2. After adjusting the final volume to 20 μl with distilled water, and PCR reaction was performed on a VeritiTM 96-well Thermal Cycler (Applied Biosystems, Carlsbad, California, USA).
Amplification of partial ompA/B and sca4
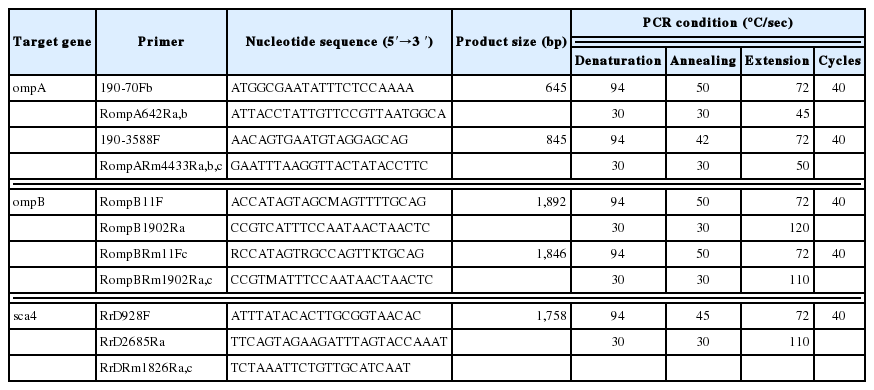
To amplify partial ompA, ompB, and sca4 genes from SFG Rickettsia positive DNA samples, nPCR was performed. Primers are listed in Table 1.
Sequencing analysis
To identify Rickettsia species by sequencing, we used ompA primers (Table 1). Sequencing was performed by Genotech Co. Ltd. (Daejeon, Korea). To acquire partial ompA nucleotide sequences, all samples were sequenced in duplicates. Sequence analyses were performed with MegAlign software (DNAStar, Madison, Wisconsin, USA).
RESULTS
Tick collection
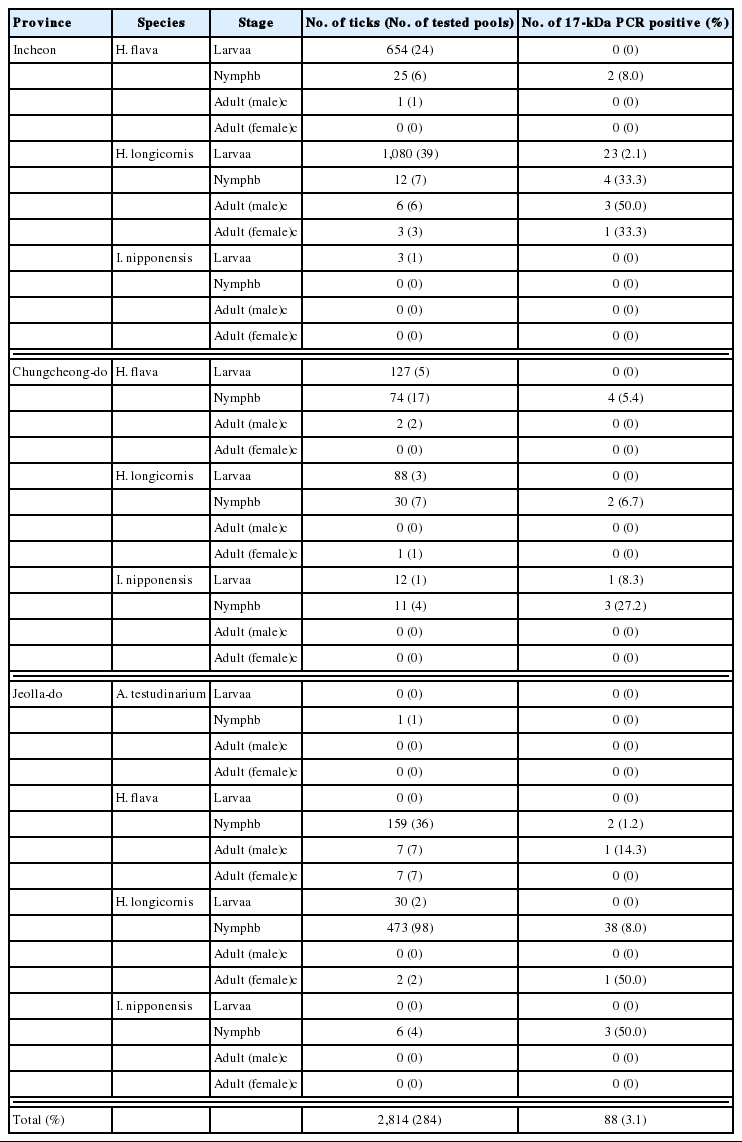
A total of 2,814 ticks were collected from 3 provinces in the Republic of Korea in May 2009, including 1,056 H. flava, 1,725 H. longicornis, 32 I. nipponensis, and one A. testudinarium. These ticks consisted of 1,994 (70.8%) larvae, 791 (28.1%) nymphs, 16 (0.5%) adult males, and 13 (0.4%) adult females. Dominant species were H. longicornis (61.3%) followed by H. flava (37.5%) and I. nipponensis (Table 2).
Amplification and sequencing for rickettsial agent identifications
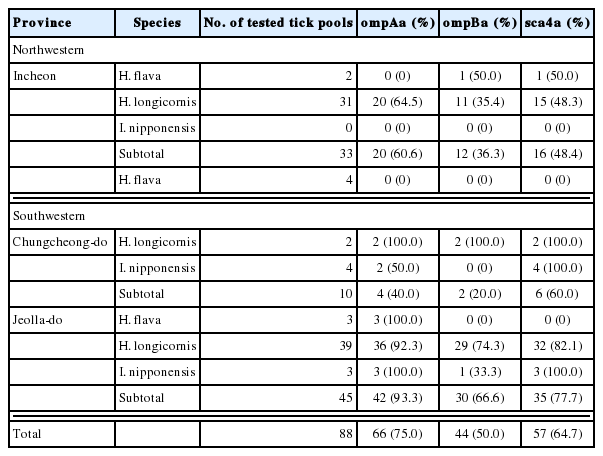
nPCR screening of 284 tick pools identified 88 (30.9%) positive samples using rickettsial 17-kDa antigen gene-specific primers. These nPCR positive samples were used for nPCR amplification of ompA, ompB, and sca4 genes. nPCR results showed that 66 (75.0%), 44 (50.0%), and 57 (64.7%) samples were positive for ompA, ompB, and sca4, respectively (Table 3).
Subsequently, we randomly selected 30 nPCR positive samples (10 from northwestern and 20 from southwestern province) for ompA gene to performed sequencing analysis.
Sequences of ompA from 2 H. flava pool samples collected from Jeolla-do shared 97.8–99.1% similarities with those of R. heilongjiangensis. Most of 27 H. longicornis pools shared 97.8–98.8% sequence similarities with R. heilongjiangensis while 1 H. longicornis tick pool shared 99.9% sequence similarity with R. monacensis (Fig. 2). R. monacensis was first detected in I. nipponensis collected from Jeolla-do, Gyeonggi-do, and Gangwon-do [22,23]. Interestingly, R. monacensis was first detected in H. longicornis collected from Incheon metropolitan city of a northwestern province.
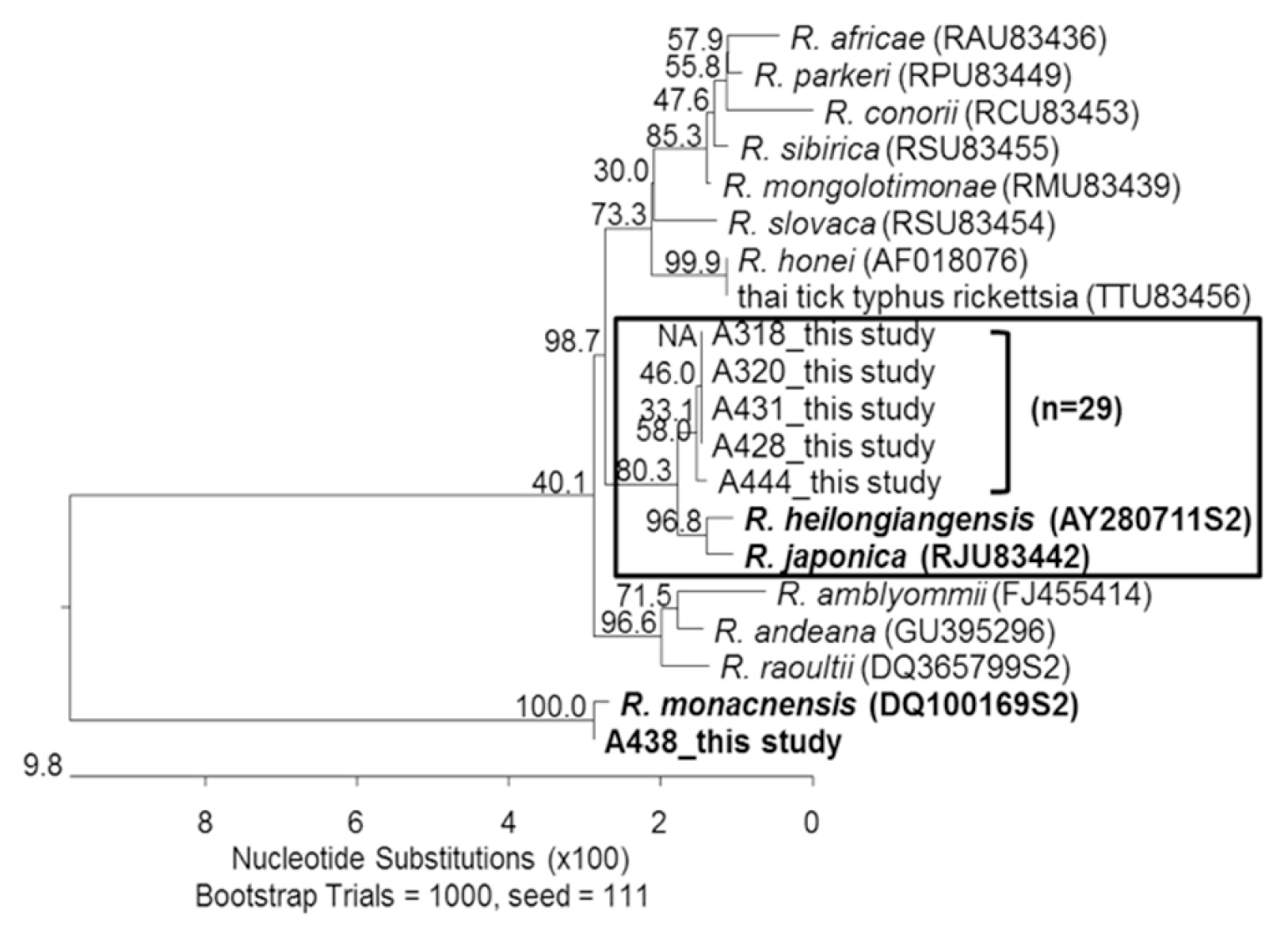
Phylogenic tree representing phylogenetic relationships between partial ompA sequence of various rickettsial strains and 625 bp of ompA product amplified from 30 selected DNA samples. The phylogenetic tree was constructed using MegAlign software and Bootstrap analysis was performed with 1,000 replicates.
DISCUSSION
First cases of Far East spotted fever (FESF) caused by R. heilongjiangensis have been reported in Russia and China [24]. Rickettsiae from ticks collected in Korea in 2003 [13] showed high sequence similarities with R. japonica YH (GenBank accession number: AP011533). R. japonica was detected in Korean human sera in 2004 and 2005 [14,16].
Although this study was limited to a short period of 5 months, most tick-related pathogens such as tick-borne pathogens found in other Korean studies [19,26,27] were commonly detected in Southern provinces such as Jeolla-do and Chungcheong-do that were also included in the present study.
To obtain more data on the distribution of rickettsiae, we investigated species of Rickettsia from ticks in 2 provinces of Republic of Korea. In particular, the number of ticks collected from Incheon metropolitan city was more than that collected from other regions and H. longicornis predominated. Its number collected from Incheon metropolitan city was twice of that collected from Jeolla-do and 9 times of that collected from Chungcheong-do.
Incheon metropolitan city is located in the northwestern part of Seoul. It is the third largest city after Seoul and Busan in Republic of Korea. Interestingly, the 8 areas of Incheon-si where ticks were collected were mostly flat areas not exceeding 100 m in height with a humid subtropical climate [28,29]. This environment is a suitable place for the survival of tick vectors and the area with low grass height may be advantageous for human and vector contact. This shows the potential that human infections can be caused by ticks in urban areas. It also reminds us that we need to continuously monitor geographical changes of vector distribution and disease incidence.
In summary, we used nucleic acids and found that rickettsial agents from Ixodid ticks collected from northwestern and southwestern provinces of the Republic of Korea were closely related to R. heilongjiangensis, R. japonica, and R. monacensis.
ACKNOWLEDGMENTS
Yeon-Joo Choi and Ju Jiang contributed equally. Funding for portions of this work was provided by the Armed Forces Health Surveillance Branch-Global Emerging Infections Surveillance and Response System (AFHSB-GEIS), Silver Spring, Maryland, USA. The views expressed in this article are those of the authors do not reflect the official policy or position of the Department of the Navy, the Department of the Army, the Department of Defense, nor the US Government.
Notes
CONFLICT OF INTEREST
The authors declare that they have no conflict of interest.