Abstract
Until now, 3 Metagonimus spp. (M. yokogawai, M. takahashii, and M. miyatai) causing human metagonimiasis have been reported in Korea. In this study, we investigated the possible presence of Metagonimus spp. other than these three species using human fecal samples from an endemic area in Korea. DNA was extracted from Metagonimus egg-positive fecal samples collected from residents of Gurye-gun, Jeollanam-do. A total of 21 representative mitochondrial cytochrome c oxidase subunit I sequences were obtained by PCR and cloning, and sequencing. Phylogenetic analysis revealed 1 cluster corresponding to M. yokogawai (n = 10) and two additional distinct clusters corresponding to M. kogai (n = 8) and M. saitoi (n = 3), which were proposed as new species in Japan in 2022. Pairwise cytochrome c oxidase subunit I distances were low for M. yokogawai and M. kogai (mean Kimura 2-parameter: 0.005–0.006), whereas M. saitoi showed higher Korea–Japan values (~0.029), a pattern consistent with geographic structuring. In conclusion, we provide the first molecular evidence for the occurrence of M. kogai and M. saitoi in human fecal samples in Korea. Further confirmation using adult morphology, additional nuclear markers, and ecological surveys are needed to clarify metagonimiasis transmission in the Seomjin-gang (river) basin.
-
Key words: Metagonimus, cytochrome c oxidase subunit I, metagonimiasis, human fecal samples
Introduction
The genus
Metagonimus includes a group of intestinal trematodes that are widely distributed in Asia, many of which are zoonotic parasites transmitted through the consumption of raw or undercooked freshwater fish [
1].
Metagonimus spp. are the causative agents of metagonimiasis, a foodborne parasitic disease with significant public health implications in endemic areas such as Korea, Japan, China, and Russia. Globally, more than 50 million cases of intestinal fluke infections, including those caused by
Metagonimus spp., have been recorded [
2]. While most infections are asymptomatic, heavy or repeated infections may cause gastrointestinal symptoms, such as diarrhea, abdominal pain, and indigestion. The ectopic migration of heterophyid flukes to extraintestinal organs such as the heart and brain has rarely been reported [
3]. Human metagonimiasis is typically diagnosed via the microscopic examination of eggs in fecal samples, commonly using the Kato–Katz and formalin-ether sedimentation methods [
4]; however, such methods are not sufficient for precise species-level discrimination.
Since the first description of
M. yokogawai in Taiwan in 1912 [
5], the diversity within this genus has become increasingly evident. Two additional species (
M. ovatus and
M. minutus) have been reported in Taiwan [
6,
7] and at least 5 species have been reported in Japan (
M. takahashii,
M. katsuradai,
M. otsurui,
M. miyatai, and
M. hakubaensis) [
8-
12];
M. suifunensis and
M. pusillus have been reported in Russia [
13,
14]. More recently, 4 new cryptic species—
M. saitoi,
M. kogai,
M. shimazui, and
M. kinoi—were proposed as new species in Japan, despite their morphological similarities to previously known species [
15]. These species were primarily characterized based on metacercariae recovered from freshwater fish, without direct evidence of human infection at the time of their initial description.
Molecular approaches are essential for delineating the species boundaries and understanding the epidemiology of
Metagonimus infections [
15]. The above findings highlight the utility of combining molecular markers, such as the mitochondrial cytochrome
c oxidase subunit I (COI) gene, nuclear 28S rDNA, and internal transcribed spacer (ITS) regions, in DNA barcoding approaches to achieve accurate species identification and resolve cryptic diversity within the genus [
15-
17]. Among these markers, COI has been recognized as the most effective one in resolving species-level differences, particularly among closely related or cryptic species within
Metagonimus and other trematodes [
15,
18,
19]. In Korea, only 3 species,
M. yokogawai,
M. takahashii, and
M. miyatai, have been reported to date [
20].
In this study, we aimed to investigate the species composition of
Metagonimus in Gurye-gun (county), along the Seomjin-gang (river), Jeollanam-do (province), an area with a persistently high prevalence of intestinal fluke infections [
21-
24], by conducting COI-based molecular analyses of human fecal samples. According to the Intestinal Parasite Infection Survey Project, the prevalence of
Metagonimus spp. infection in Gurye-gun has been reported to be ≥3.0%, which is notably higher than the national average level of 1.7% [
21-
24]. As low egg counts in samples often lead to insufficient PCR sensitivity for direct sequencing, DNA was extracted from pooled samples, and molecular cloning was employed to isolate individual sequences. Using this approach, we aimed to provide COI-based molecular evidence for the possible occurrence of previously unrecorded
Metagonimus spp. in Korea, rather than categorical identification.
Methods
Ethics statement
Ethical approval was waived because this study was conducted to evaluate public welfare through a fact-finding survey (Infectious Disease Control and Prevention Act, Article 17(1)).
Sample collection
Fecal samples were collected from residents of Gurye-gun, Jeollanam-do, Korea, as part of the 2024 National Intestinal Parasite Survey conducted by the Korea Disease Control and Prevention Agency. Gurye-gun was selected because of consistently higher prevalence of
Metagonimus spp. infections over the past 5 years (2020–2024) than in the other regions [
21-
24].
All fecal specimens were stored at 4°C until further processing. Samples were prepared using the ParaEgg Kit (LM Science) [
25], which employs a simplified water–ether concentration method. Among the 124 samples microscopically confirmed as positive for intestinal parasite infection (including cases of mixed infection), 24 were excluded because of co-infection with other intestinal helminths, such as
Clonorchis sinensis and
Trichuris trichiura. They were excluded to prevent amplification of non-target species during molecular cloning, which could otherwise lead to misidentification of
Metagonimus-specific sequence data. Consequently, only 100 samples confirmed to be infected with
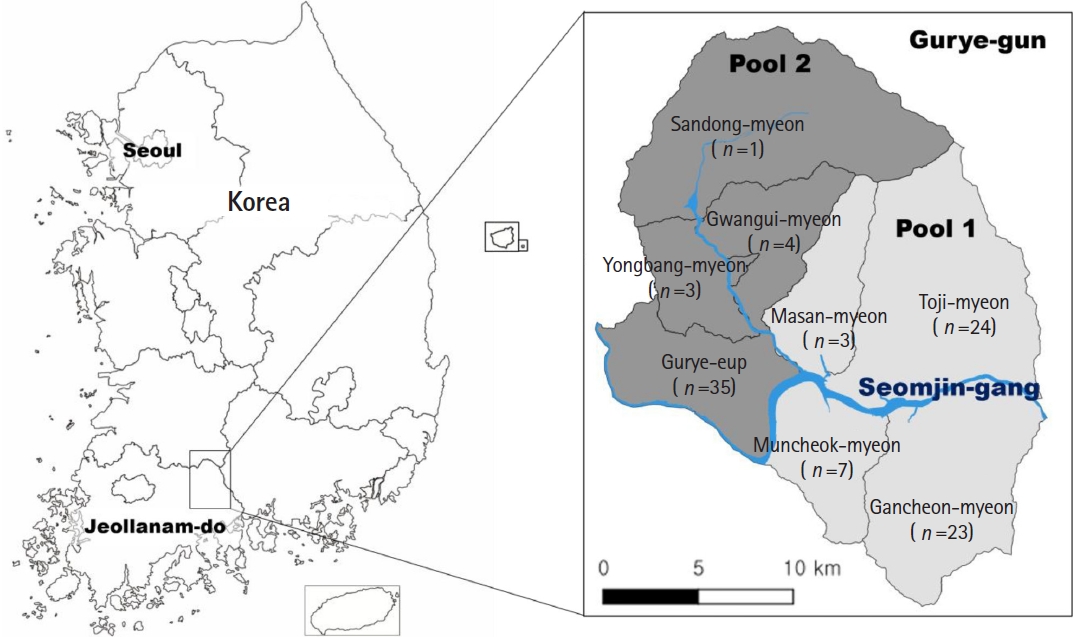
Metagonimus sp. were selected for molecular analysis. To facilitate DNA extraction and ensure representative coverage across the study area, 100 samples were divided into 2 pools according to administrative subdivisions (
Fig. 1).
DNA was extracted separately from each of the 2 pooled samples using the DNeasy 96 PowerSoil Pro Kit (Qiagen) on the QIAcube HT automated nucleic acid extractor (Qiagen). The manufacturer’s protocol was modified to include a bead-beating step to enhance DNA yield [
26].
For DNA barcoding, the COI gene was amplified using the universal primer set JB3/JB4.5 [
27], with the following sequences: JB3(F) 5’-TTTTTTGGGCATCCTGAGGTTTAT-3’, JB4.5(R) 5’-TAAAGAAAGAACATAATGAAAATG-3’.
PCRs were carried out using AccuPower PCR PreMix (Bioneer) in a 20-μl total volume, containing 5 μl of template DNA and 0.5 μM of each primer. The PCR conditions were as follows: initial denaturation at 95°C for 5 min; followed by 40 cycles at 95°C for 20 sec, 48°C for 20 sec, and 72°C for 1 min; and a final extension at 72°C for 5 min.
The amplicons were separated using electrophoresis on a 1.5% agarose gel and purified using the QIAquick Gel Extraction Kit (Qiagen) according to the manufacturer’s instructions.
Cloning and sequencing
The purified PCR products were cloned into the pCR2.1-TOPO vector using the TOPO TA Cloning Kit (Invitrogen) according to the manufacturer’s protocol. The ligation mixtures were transformed into Stellar Competent Cells (Takara) via heat shock at 42°C for 90 sec, followed by immediate incubation on ice for 2 min.
The transformed cells were spread on LB agar plates supplemented with ampicillin (100 μg/ml) and X-gal (20 mg/ml) and incubated at 37°C for 16–18 h. A total of 60 white colonies (30 colonies per plate) were randomly selected from 2 plates to ensure a balance between representativeness and sequencing efficiency. The selected colonies were individually suspended in 50 μl of LB broth. After 1 min of boiling, the samples were centrifuged, and the supernatant was used as template DNA for PCR, which was performed under the same conditions as those described in Section ‘DNA extraction and amplification.’
The final PCR products were sequenced by a commercial service provider (SolGent).
Phylogenetic analysis
The obtained COI sequences were aligned using the ClustalW algorithm implemented in MEGA version 12. Phylogenetic trees were constructed using the maximum likelihood method with the Tamura–Nei model. The robustness of the phylogenetic groupings was evaluated using 1,000 bootstrap replicates.
To assess genetic distances within species, pairwise genetic distances were calculated using the Kimura 2-parameter (K2P) model.
Results
COI sequence acquisition
A total of 100 Metagonimus spp.–positive fecal samples were divided into 2 pools based on their geographic location in Gurye-gun. From these 2 pooled DNA samples, 60 white colonies were selected after TA cloning and subjected to COI gene sequencing. Among them, 21 high-quality sequences with trimmed lengths ranging from 392 to 397 bp were successfully obtained, with 7 and 14 colonies from Pools 1 and 2, respectively. All sequences were submitted to GenBank (accession No. PV876432– PV876452) and were included in subsequent phylogenetic analysis.
Sequence analysis and species identification
All 21 sequences were subjected to BLAST search and phylogenetic analysis. The BLAST search results showed high similarity (≥97%) of the acquired sequences to the reference sequences of
M. yokogawai,
M. kogai, and
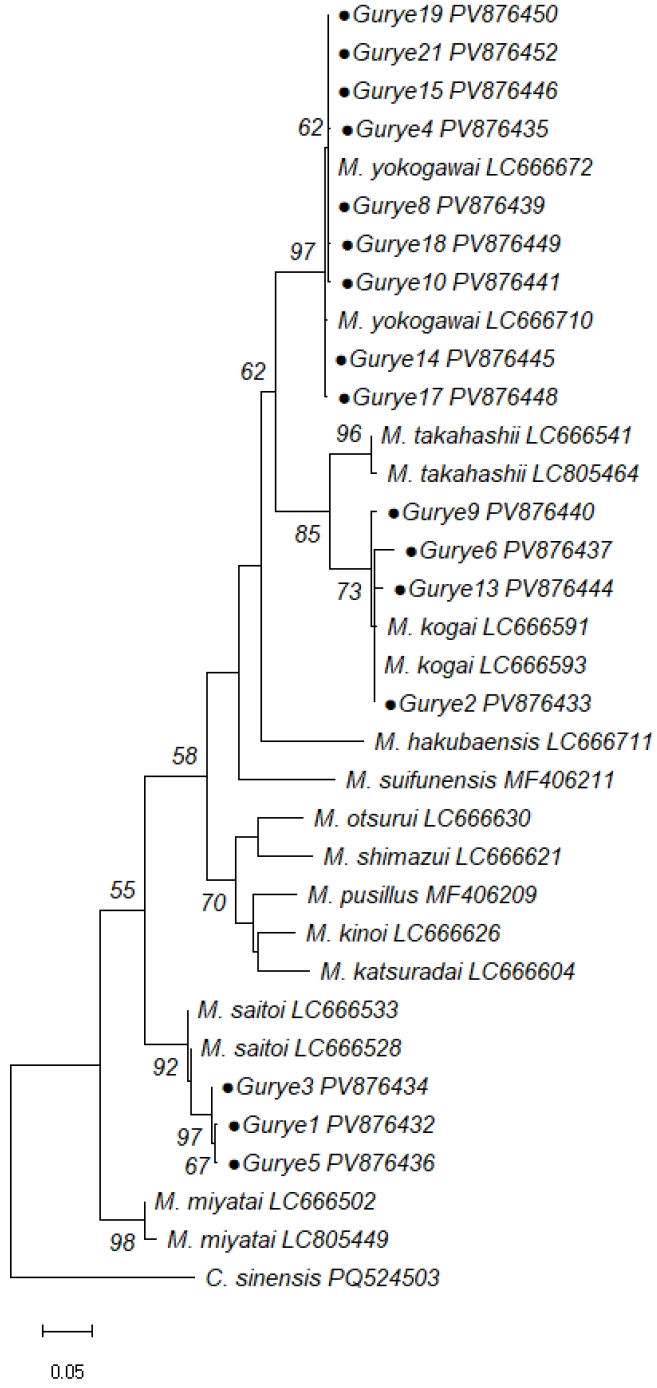
M. saitoi. The phylogenetic analysis using MEGA12 based on the maximum likelihood method with the Tamura–Nei model revealed that all sequences clustered with their respective reference species (
Fig. 2). These findings provide molecular evidence of the presence of 3 distinct
Metagonimus spp. in Korea.
Genetic distances (K2P) between the sequences obtained in this study and the Japanese reference sequences were calculated to assess intraspecific and interspecific variation. As shown in
Table 1, the mean intraspecific genetic divergence between the Korean and Japanese isolates was 0.005 for
M. yokogawai and 0.006 for
M. kogai, indicating a high degree of sequence similarity. In
M. saitoi, a slightly higher mean intraspecific genetic divergence was observed (0.029). Additionally, the interspecific genetic divergence values calculated among the Gurye-gun samples were 0.126 between
M. yokogawai and
M. kogai, 0.152 between
M. yokogawai and
M. saitoi, and 0.197 between
M. kogai and
M. saitoi, confirming a clear genetic differentiation among these 3 species.
Discussion
In this study, we conducted a molecular analysis of COI gene sequences obtained from
Metagonimus-positive fecal samples collected from residents of Gurye-gun, Jeollanam-do, Korea. A total of 21 sequences were successfully obtained and identified as those of
M. yokogawai (
n=10),
M. kogai (
n=8), and
M. saitoi (
n=3), based on their high similarity (≥97%) to previously reported sequences (
Fig. 2). Notably, the occurrence of
M. kogai and
M. saitoi, which were proposed as new species in Japan in 2022 [
15], in Korea was supported by COI-based molecular evidence from human fecal samples.
The initial Japanese descriptions of
M. kogai and
M. saitoi were based on morphological and molecular analyses of metacercariae collected from freshwater fish and adult worms recovered from experimentally infected animals [
15]. In contrast, our findings, derived from analysis of
Metagonimus eggs in human fecal samples, provided clear evidence that humans can serve as definitive hosts for these species, highlighting their importance in public health.
To date, 3
Metagonimus spp. (
M. yokogawai,
M. takahashii, and
M. miyatai) have been reported in Korea.
M. yokogawai is predominantly found along the southern and eastern riversides, whereas
M. takahashii and
M. miyatai have been identified mainly in inland regions, particularly in the upper reaches of large rivers [
20,
28-
30]. Notably, all 3 species were previously reported to be isolated from freshwater fish in Hadong-gun (county) within the Seomjin-gang (river) basin, which is downstream of Gurye-gun [
28]. However, in the present study,
M. takahashii and
M. miyatai were not detected in the samples collected from Gurye-gun. This discrepancy in the findings may reflect local differences in intermediate host availability or ecological conditions between the upper (Gurye-gun) and lower (Hadong-gun) reaches of the Seomjin-gang Basin, as well as potential differences arising from human versus fish sampling.
According to Nakao et al. [
15],
M. kogai and
M. saitoi are classified within the
M. takahashii and
M. miyatai complexes, respectively, owing to their considerable morphological similarities. This classification implies that earlier reports, based solely on morphological characteristics, may have overlooked the presence of these newly described species in Korea. Furthermore, a phylogenetic analysis using the 28S rRNA gene placed
M. kogai and
M. takahashii within the same genetic cluster [
15], further highlighting the limitations of morphology-based taxonomy. In this milieu, we examined within- and between-region COI divergence to interpret population structure.
In this study, the intraspecies genetics of M. saitoi within the Gurye-gun region of Korea was assessed by comparing the COI sequences of M. saitoi obtained from fecal samples. The mean pairwise K2P genetic distance between Korean M. saitoi sequences was 0.005, indicating a very low level of divergence. This finding supports their classification as a single species and suggests high genetic homogeneity within the local population. Such genetic uniformity is commonly observed in populations with limited geographical barriers and continuous gene flow.
The average intraspecific genetic distance between the Korean and Japanese
M. saitoi sequences was 0.029, which was relatively high, although it was still within or near the expected range for intraspecific variation in trematodes. This relatively high divergence raises the possibility of geographic structuring between the Korean and Japanese populations, which could be attributed to habitat fragmentation or limited host dispersal commonly observed among freshwater trematodes [
31]. In contrast,
M. yokogawai and
M. kogai exhibited very low intraregional genetic distances of 0.005 and 0.006, respectively, indicating strong genetic similarity and limited genetic differentiation between the Korean and Japanese populations. This finding suggests either recent or ongoing gene flow between Korean and Japanese populations or the presence of similar ecological niches and host associations that constrain genetic divergence across these regions [
32].
A limitation of this study was that the pooled fecal samples were collected from a single endemic region (Gurye-gun). As pooling and the morphological indistinguishability of eggs precluded individual-level resolution, we could not assess within-host co-infection involving multiple Metagonimus spp., warranting future individual-based sampling. In addition, future studies should involve individual-based sampling and perform multi-basin comparisons across the Yeongsan-gang, Nakdong-gang, and Seomjin-gang (rivers) in Korea.
Other limitations include reliance of our study on a single mitochondrial locus (COI), lack of adult worm morphology, and absence of ecological data (including data on intermediate hosts and transmission foci). In our future research, we will recover adults for integrative morphology, identify intermediate hosts and local transmission foci within the Seomjin-gang (river) system. We will incorporate nuclear markers (28S rDNA and ITS) to test COI-based inferences and evaluate population structure, including potential Korea–Japan differentiation in M. saitoi.
From a public-health perspective, coupling COI assays with microscopy may improve species-level attribution in endemic basins; however, confirmatory multi-gene barcoding and adult morphology remain essential before any taxonomic revision. Our molecular evidence updates the working species spectrum for Korea and provides a framework for subsequent morphological and ecological investigations.
Notes
-
Author contributions
Conceptualization: Lee E, Lee HI, Ju JW, Lee MR. Investigation: Lee. Resources: Lee YJ, Baek SO. Software: Choi JH. Supervision: Kim TY. Writing – original draft: Lee E. Writing – review & editing: Lee E, Kim TY.
-
Conflict of interest
The authors have no conflicts of interest to declare.
-
Funding
This study was supported by a grant (6332-305-210-13, 2025) from the Korea Disease Control and Prevention Agency (KDCA), Republic of Korea.
-
Acknowledgments
We thank the local officials of Gurye-gun, including Mi Suk Park and Mun Young Jang, for their support in sample collection and Ji Hoon Ha, Ki Min Kim, and Kyung Hwa Pung of the Korea Association of Health Promotion for their assistance in diagnosis.
Fig. 1.Survey areas for detecting Metagonimus eggs in Gurye-gun, Jeollanam-do, Korea. Subregions shaded light gray correspond to Pool 1, whereas those shaded dark gray represent Pool 2. The number of positives included in each pool is shown under the name of town (eup) and township (myeon). Blue line indicates streams of Seomjin-gang (river).
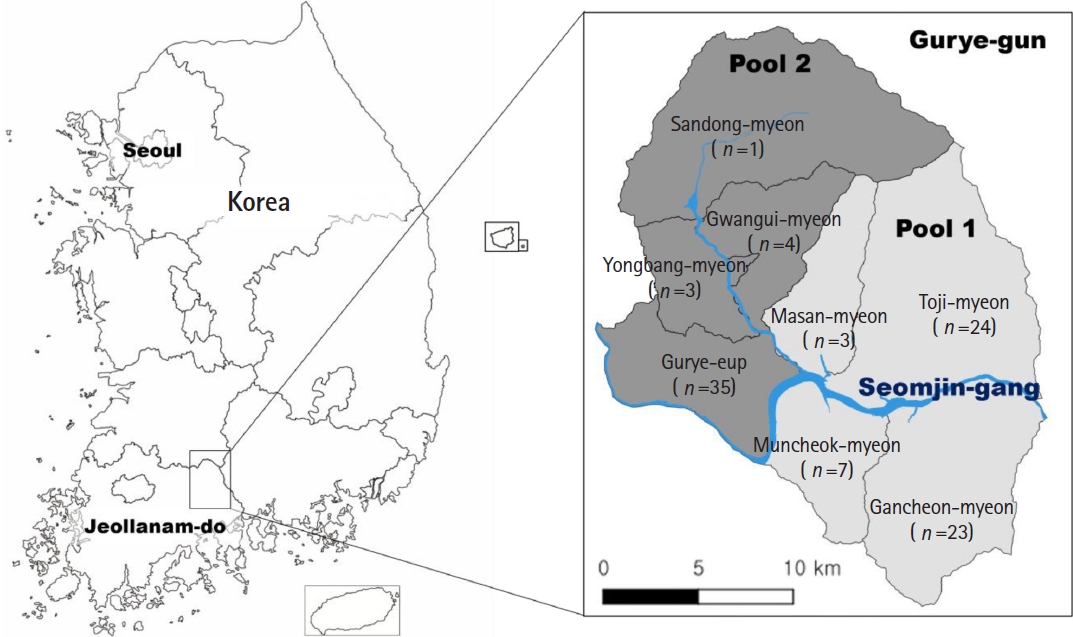
Fig. 2.Maximum likelihood phylogenetic tree based on cytochrome c oxidase subunit I sequences of Metagonimus spp. identified in this study and reference sequences. Identical sequences were collapsed for clarity; Gurye15 represents both Gurye15 and Gurye20; Gurye2 represents Gurye2, Gurye7, Gurye11, Gurye12, and Gurye6. Bootstrap percentages are shown on representative nodes.
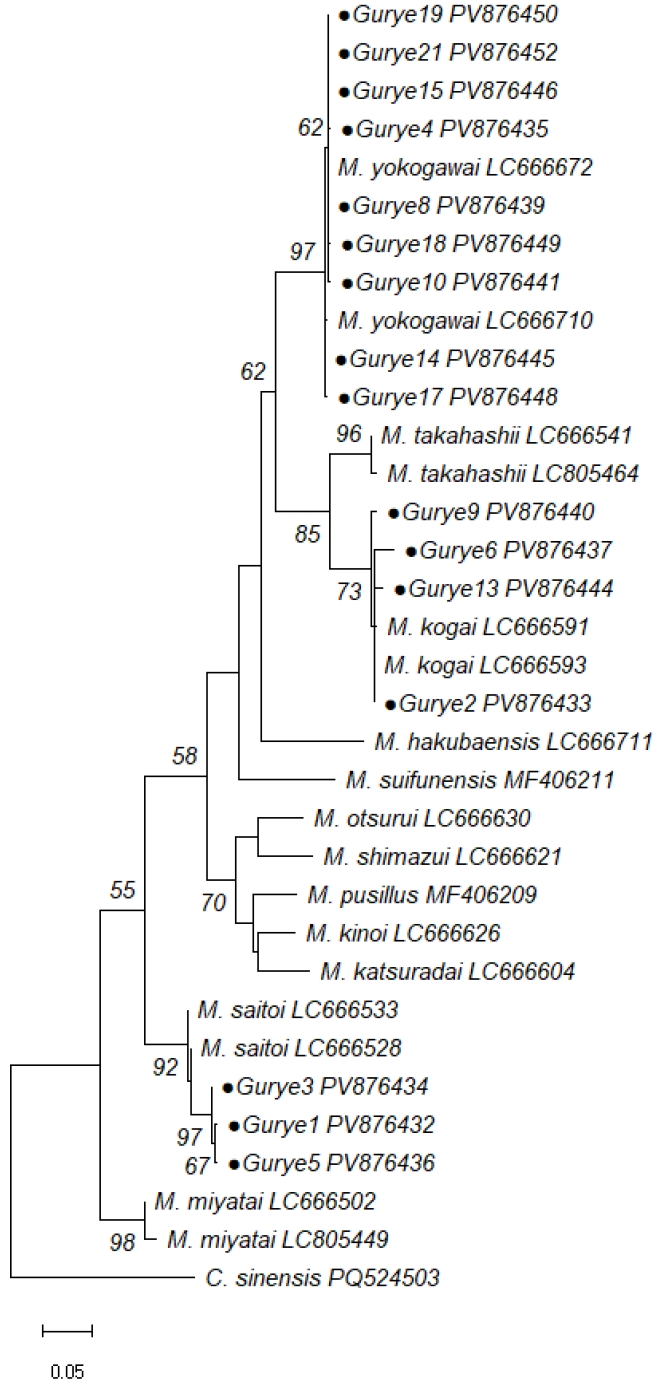
Table 1.Intraspecific and interspecific genetic divergence (Kimura 2-parameter) of Metagonimus species based on cytochrome c oxidase subunit I sequences
Table 1.
|
Species |
Intraspecific (Korea–Japan) |
Intraspecific (within Gurye-gun) |
Interspecific (within Gurye-gun) |
|
My |
Mk |
Ms |
|
My (n=10) |
0.005 (0.000–0.011) |
0.006 (0.000–0.010) |
NA |
0.126 |
0.152 |
|
Mk (n=8) |
0.006 (0.000–0.022) |
0.009 (0.000–0.028) |
0.126 |
NA |
0.197 |
|
Ms (n=3) |
0.029 (0.022–0.034) |
0.005 (0.005–0.005) |
0.152 |
0.197 |
NA |
References
- 1. Chai JY, Hong ST, Choi MH, Shin EH, Bae YM. Clinical parasitology. Seoul National University Press; 2011.
- 2. Fürst T, Keiser J, Utzinger J. Global burden of human food-borne trematodiasis: a systematic review and meta-analysis. Lancet Infect Dis 2012;12:210-21. https://doi.org/10.1016/S1473-3099(11)70294-8
- 3. Chai JY, Jung BK. Fishborne zoonotic heterophyid infections: an update. Food Waterborne Parasitol 2017;8:33-63. https://doi.org/10.1016/j.fawpar.2017.09.001
- 4. Montresor A, Crompton DW, Hall A, et al. Guidelines for the evaluation of soil-transmitted helminthiasis and schistosomiasis at community level: a guide for managers of control programmes. World Health Organization; 1998.
- 5. Katsurada F. Heterophyes in Japan. Okayama Igakkai Zasshi 1912;268:373-80.
- 6. Yokogawa S. On a new parasite Metagonimus yokogawai with a trout, Plecoglossus altivelis (Temminck), as intermediate host: a new genus established. Tokyo Igakkai Zasshi 1913;72:158-79.
- 7. Katsuta I. Studies on trematodes whose second intermediate hosts are fishes from the brackish waters of Formosa. Taiwan Igakkai Zasshi 1932;31:26-39.
- 8. Takahashi S. On the life-history of Metagonimus yokogawai, a new species of Metagonimus and Exorchis major. Okayama Igakkai Zasshi 1929;42:2687-755.
- 9. Izumi M. Studies concerning a new species of Metagonimus and its life cycle. Kitasato Arch Exp Med 1935;12:365-84.
- 10. Saito S, Shimizu T. A new trematode, Metagonimus otsurui sp. nov. from the fresh-water fishes (Trematoda: Heterophyidae). Jpn J Parasitol 1968;17:167-74.
- 11. Saito S, Chai JY, Kim KH, Lee SH, Rim HJ. Metagonimus miyatai sp. nov. (Digenea: Heterophyidae), a new intestinal trematode transmitted by freshwater fishes in Japan and Korea. Korean J Parasitol 1997;35:223-32. https://doi.org/10.3347/kjp.1997.35.4.223
- 12. Shimazu T. Metagonimus hakubaensis sp. n. (Digenea, Heterophyidae) from Nagano, Japan: morphology and life cycle. Bull Natl Sci Mus Ser Zool 1999;25:87-99.
- 13. Shumenko PG, Tatonova YV, Besprozvannykh V. Metagonimus suifunensis sp. n. (Trematoda: Heterophyidae) from the Russian Southern Far East: morphology, life cycle, and molecular data. Parasitol Int 2017;66:982-91. https://doi.org/10.1016/j.parint.2016.11.002
- 14. Tatonova YV, Shumenko PG, Besprozvannykh VV. Description of Metagonimus pusillus sp. nov.(Trematoda: Heterophyidae): phylogenetic relationships within the genus. J Helminthol 2018;92:703-12. https://doi.org/10.1017/S0022149X17001146
- 15. Nakao M, Ishikawa T, Hibino Y, et al. Resolution of cryptic species complexes within the genus Metagonimus (Trematoda: Heterophyidae) in Japan, with descriptions of four new species. Parasitol Int 2022;90:102605. https://doi.org/10.1016/j.parint.2022.102605
- 16. Lee SU, Huh S, Sohn WM, Chai JY. Sequence comparisons of 28S ribosomal DNA and mitochondrial cytochrome c oxidase subunit I of Metagonimus yokogawai, M. takahashii and M. miyatai. Korean J Parasitol 2004;42:129-35. https://doi.org/10.3347/kjp.2004.42.3.129
- 17. Pornruseetairatn S, Kino H, Shimazu T, et al. A molecular phylogeny of Asian species of the genus Metagonimus (Digenea)—small intestinal flukes—based on representative Japanese populations. Parasitol Res 2016;115:1123-30. https://doi.org/10.1007/s00436-015-4843-y
- 18. Bowles J, Blair D, McManus DP. A molecular phylogeny of the human schistosomes. Mol Phylogenet Evol 1995;4:103-9. https://doi.org/10.1006/mpev.1995.1011
- 19. Vilas R, Criscione CD, Blouin MS. A comparison between mitochondrial DNA and the ribosomal internal transcribed regions in prospecting for cryptic species of platyhelminth parasites. Parasitology 2005;131:839-46. https://doi.org/10.1017/S0031182005008437
- 20. Chai JY, Lee SH. Food-borne intestinal trematode infections in the Republic of Korea. Parasitol Int 2002;51:129-54. https://doi.org/10.1016/s1383-5769(02)00008-9
- 21. Lee MR, Ju JW, Baek SO, Lee YJ, Lee HI. Infection status of intestinal helminths in 2021. Public Health Wkly Rep 2022;15:1773-83.
- 22. Lee MR, Ju JW, Baek SO, Lee YJ, Lee HI. Infection status of intestinal helminths in 2022. Public Health Wkly Rep 2023;16:1131-40. https://doi.org/10.56786/PHWR.2023.16.32.2
- 23. Lee MR, Ju JW, Baek SO, et al. Infection status of intestinal helminths in 2023. Public Health Wkly Rep 2024;17:1227-39. https://doi.org/10.56786/PHWR.2024.17.28.2
- 24. Kim TY, Lee MR, Ju JW, et al. Survey on intestinal helminth infections in the endemic regions, 2024. Public Health Wkly Rep 2025;18:707-19. https://doi.org/10.56786/PHWR.2025.18.19.1
- 25. Lee E, Back SO, Lee YJ, et al. A novel kit for enrichment of fecal helminth eggs. Parasites Hosts Dis 2024;62:323-9. https://doi.org/10.3347/PHD.24021
- 26. Srirungruang S, Mahajindawong B, Nimitpanya P, et al. Comparative study of DNA extraction methods for the PCR detection of intestinal parasites in human stool samples. Diagnostics (Basel) 2022;12:2588. https://doi.org/10.3390/diagnostics12112588
- 27. Bowles J, Blair D, McManus DP. Genetic variants within the genus Echinococcus identified by mitochondrial DNA sequencing. Mol Biochem Parasitol 1992;54:165-73. https://doi.org/10.1016/0166-6851(92)90109-w
- 28. Chai JY, Sohn WM, Kim MH, Hong ST, Lee SH. Three morphological types of the genus Metagonimus encysted in the dace, Tribolodon taczanowskii, caught from the Sumjin River. Korean J Parasitol 1991;29:217-25. https://doi.org/10.3347/kjp.1991.29.3.217
- 29. Chai JY, Huh S, Yu JR, et al. An epidemiological study of metagonimiasis along the upper reaches of the Namhan River. Korean J Parasitol 1993;31:99-108. https://doi.org/10.3347/kjp.1993.31.2.99
- 30. Ahn YK. Intestinal flukes of genus Metagonimus and their second intermediate hosts in Kangwon-do. Korean J Parasitol 1993;31:331-40. https://doi.org/10.3347/kjp.1993.31.4.331
- 31. Santacruz A, Ornelas‐García CP, Pérez‐Ponce de León G. Incipient genetic divergence or cryptic speciation? Procamallanus (Nematoda) in freshwater fishes (Astyanax). Zool Scr 2020;49:768-78. https://doi.org/10.1111/zsc.12443
- 32. Blasco-Costa I, Poulin R. Host traits explain the genetic structure of parasites: a meta-analysis. Parasitology 2013;140:1316-22. https://doi.org/10.1017/S0031182013000784
 , Jong-Hun Choi
, Jong-Hun Choi , Yeong-Ju Lee
, Yeong-Ju Lee , Seon-Ok Baek
, Seon-Ok Baek , Hee-Il Lee
, Hee-Il Lee , Jung-Won Ju
, Jung-Won Ju , Myeong-Ro Lee
, Myeong-Ro Lee , Tae Yun Kim,*
, Tae Yun Kim,*