Abstract
Giardia lamblia is a protozoan parasite responsible for Giardiasis, one of the most prevalent intestinal infections worldwide. Despite its medical relevance, the molecular organization of its transcriptional apparatus remains poorly characterized. Here, I present an integrative analysis of the structural and functional features of the Giardia nucleolus and its transcription machinery. Treatment with actinomycin D induces nucleolar disorganization, confirming active rRNA transcription and nucleolar stress. Additionally, I highlight the highly divergent TATA-binding protein as a potential therapeutic target, given its essential role in transcription and its low mutation rate. Collectively, these findings provide new insights into the minimalist eukaryotic architecture of G. lamblia and identify unique molecular elements that may serve as selective antiparasitic targets.
-
Key words: Giardia lamblia, nucleolus, transcription, actinomycin D, TATA-binding protein, drug targets
Introduction
Giardia lamblia is a cosmopolitan, single-celled parasite responsible for Giardiasis, one of the most common causes of diarrheal disease globally [
1,
2]. Despite its public health relevance and status as a neglected tropical disease, the molecular biology of this disease remains underexplored [
3]. Insights into its transcriptional machinery and the previously unrecognized presence of a nucleolus have revealed potential therapeutic targets unique to this parasite [
4].
G. lamblia is an important eukaryote characterized by a highly reduced cellular architecture. Its simplified nucleolus and transcriptional machinery contain divergent molecular components that differ from those of higher eukaryotes. Understanding these unique features provides fundamental information that can support future development of antiparasitic drugs. This review summarizes current structural and functional evidence to highlight their relevance as potential drug targets.
Global Relevance and Therapeutic Challenges of G. lamblia
Giardiasis affects approximately 300 million people annually, with children under 5 being the most vulnerable [
1,
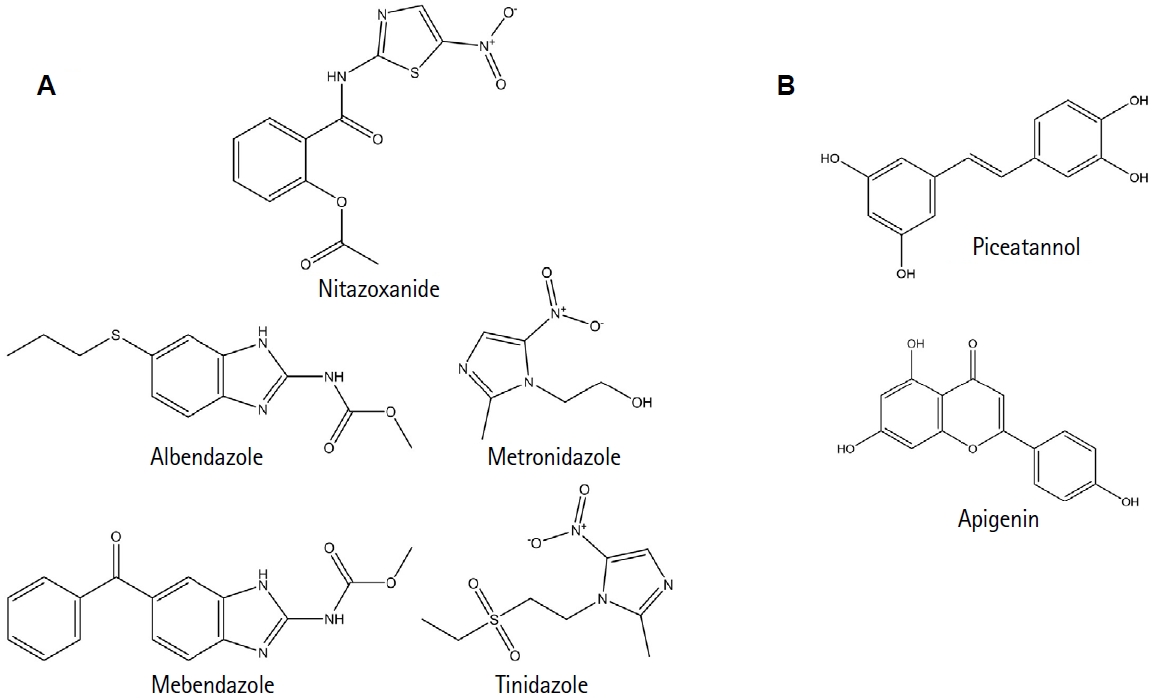
5]. Although treatable, the drugs currently used—metronidazole, tinidazole, albendazole, mebendazole, and nitazoxanide—often have adverse effects and rising reports of drug resistance (
Fig. 1A) [
6,
7]. These challenges underscore the urgent need to identify novel drug targets that exploit the parasite's unique molecular features [
8], as shown in
Fig. 1B.
Unfortunately, several of these drugs present adverse effects ranging from mild to severe, highlighting that some have been reported as genotoxic and even carcinogenic in animals. However, the potential risk in humans remains a subject of debate [
9-
11].
Discovery and Structural Characterization of the Nucleolus in G. lamblia
G. lamblia challenges traditional views of eukaryotic cell biology due to its minimalist architecture. Long considered an early-diverging eukaryote for lacking canonical organelles such as mitochondria and peroxisomes, and possessing only a rudimentary Golgi complex [
12-
14]. Recent studies suggest that its simplicity reflects a high specialization for parasitism [
14,
15]. Originally believed to lack a nucleolus, this idea was based on the homogeneous distribution of rDNA and nucleolar markers such as fibrillarin (involved in 2'-O-methylation and processing of pre-rRNA) [
16,
17]. However, the discovery of key nucleolar components like KRR1 (small subunit processome component homolog) involved in small ribosomal subunit biogenesis, CBF5 (Cajal body-specific factor 5) a pseudouridine synthase, and a set of at least 20 small nucleolar RNAs including
GlsR17 (which guide site-specific chemical modifications of rRNA), combined with ultrastructural imaging and in situ hybridization, confirmed the presence of a small but functional nucleolus (
Fig. 2A) [
4,
18-
21]. Notably, these proteins exhibit low sequence identity with their human homologs, and several of the small nucleolar RNAs lack clear orthologs in higher eukaryotes, suggesting their potential as selective therapeutic targets. This nucleolus is unusually stable throughout the cell cycle, unlike in higher eukaryotes [
20,
22]. Its reduced RNA polymerase I machinery includes only a minimal set of components required for rRNA transcription in
Giardia, reflecting the streamlined nature of its nucleolar organization [
23].
Functional Evidence of Nucleolar Activity under Transcriptional Stress
Exposure of
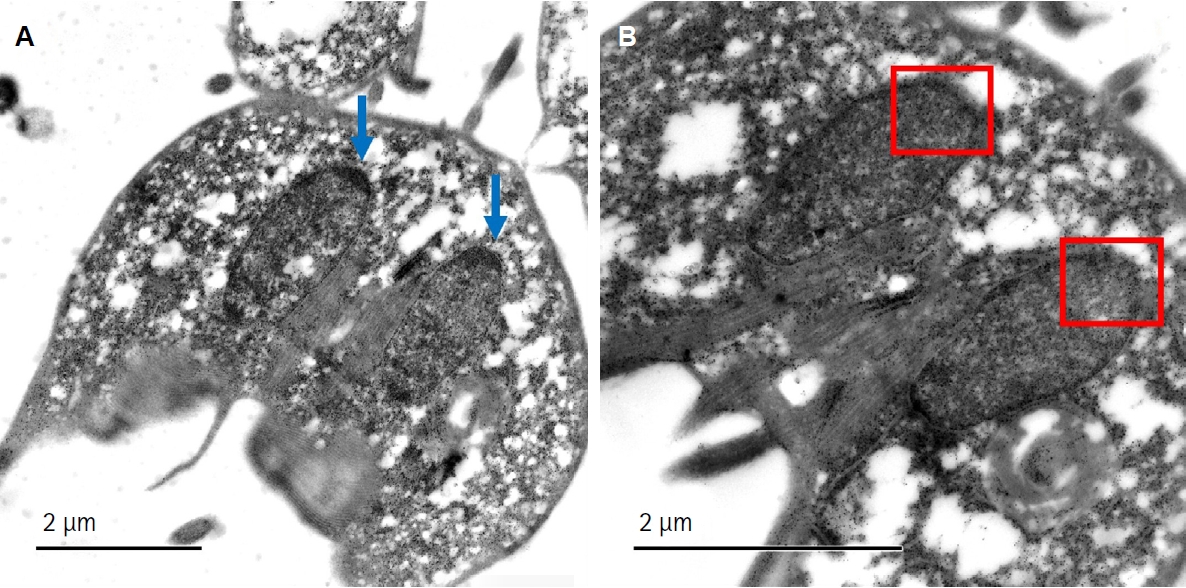
Giardia trophozoites to actinomycin D—a selective inhibitor of RNA polymerase I—leads to structural reorganization of the nucleolus. This disaggregation of nucleolar material mimics nucleolar stress observed in higher eukaryotes and indicates that
Giardia's nucleolus is actively engaged in rRNA transcription (
Fig. 2B) [
24].
A Minimalist but Essential Transcriptional Machinery
Compared to higher eukaryotes,
G. lamblia retains only 4 of the 12 basal transcription initiation factors, specifically Rrn3, A43, BRF, and C34. In addition,
Giardia encodes a highly divergent TATA-binding protein (TBP) that, although not included in the canonical set of 12 basal factors, remains essential for transcription by all 3 RNA polymerases. This TBP is remarkably unusual: it lacks 3 of the 4 conserved phenylalanine residues required for DNA binding in other eukaryotes, yet it still carries out promoter recognition and transcription initiation, highlighting the extreme structural simplification of the giardial transcription machinery [
23]. The nucleolar components discussed in this review, together with the general transcription initiation factors present in
Giardia, are summarized in
Supplementary Table 1.
Additionally, it has been reported that the sequence of the small subunit ribosomal RNA in
G. lamblia has an unusually high G-C content, close to 75%, and a length of 1,453 nucleotides—both distinctive features of prokaryotic organisms. Moreover, the small subunit ribosomal RNA of
Giardia retains the Shine-Dalgarno sequence, which is essential for ribosome binding to bacterial mRNA [
25]. This G-C richness may also enhance
Giardia's susceptibility to actinomycin D. This drug intercalates into G-C-rich DNA regions and inhibits transcription by RNA polymerase I, further underscoring the parasite's potential vulnerability due to its streamlined transcriptional machinery [
24].
The Divergent TBP as a Potential Selective Drug Target
Given the high divergence reported for the universal transcription factor TBP, and because TBP is essential for transcription by all 3 RNA polymerases, it can be postulated that this protein may be susceptible to selective therapeutic targeting [
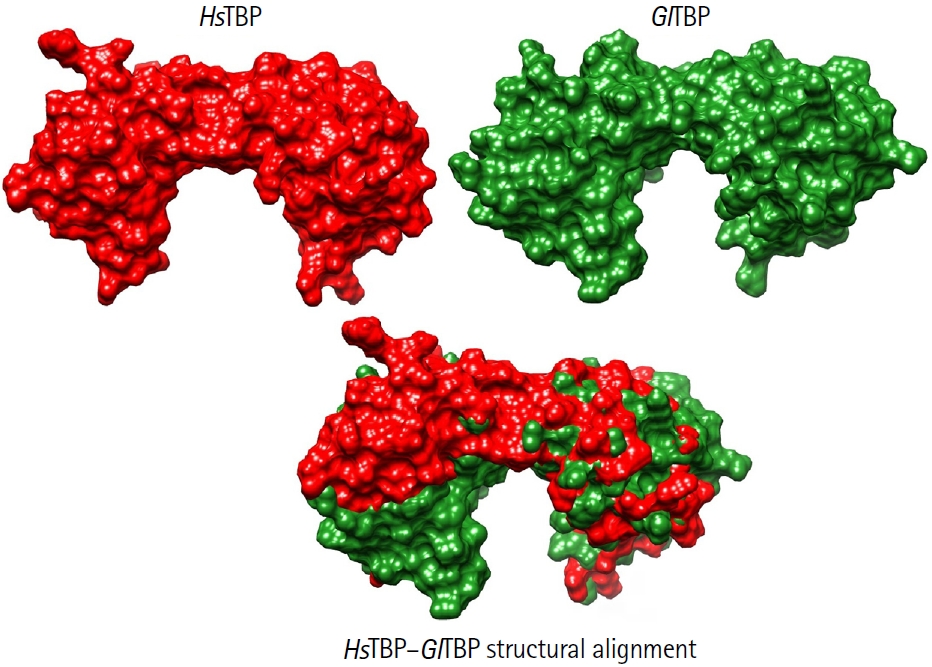
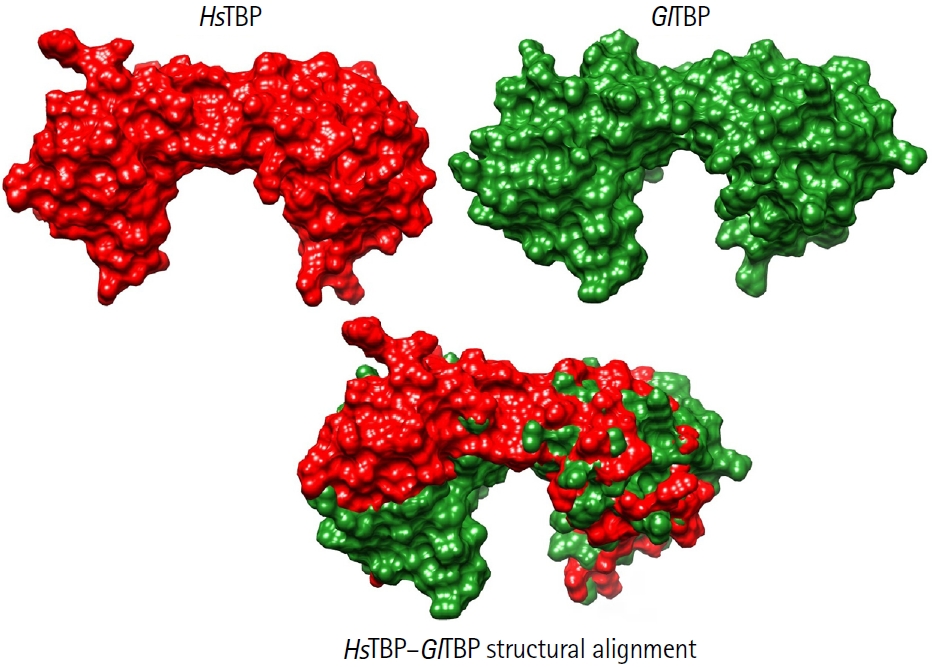
8]. This idea is supported by comparative analyses showing that TBPs from protozoan parasites exhibit markedly lower conservation than those in higher eukaryotes. As illustrated in
Fig. 3, structural comparisons between human and
Giardia TBP reveal pronounced differences in the surface topology of the protein, further highlighting its evolutionary divergence and potential suitability as a selective drug target [
26,
27]. Additionally, the different crystal structures of TBPs in complex with other transcription factors reveal that almost all proteins interact with multiple transcription factors. For example, the characteristic folding of the C-terminal domain of the TBP is a saddle-like structure; such a conformation generates 2 surfaces in the TBP. On the one hand, the convex surface interacts with various general transcription factors, which are essential for assembling the pre-initiation complex. On the other hand, the concave surface interacts with the minor groove of the DNA in the gene's promoter region, enabling transcription [
27]. Additionally, Santiago et al. [
28] reported the first systematic structural analysis of TBPs of parasites, finding many differences between the TBPs of parasites and their counterpart in higher eukaryotes, finding that the convex surface is susceptible to the use of drugs that prevent the proper assembly of the pre-initiation complex by steric hindrances, inhibiting transcription. One aspect to highlight concerning other therapeutic targets is that TBP has an extremely low mutation rate; therefore, the problem of developing resistance to drugs that target this protein would be significantly more difficult for the pathogen to achieve [
28].
Recently, in silico analyses identified compounds such as apigenin and piceatannol as potential TBP inhibitors, supporting in vitro data showing impaired parasite growth [
8] (
Fig. 1B).
Conclusion
The study of G. lamblia's nucleolus and transcription factors reveals fundamental aspects of eukaryotic biology and offers unexpected therapeutic opportunities. Targeting its minimalist yet essential molecular machinery could lead to more effective, selective antiparasitic treatments.
Notes
-
Author contributions
Conceptualization: Gaona-López C. Data curation: Gaona-López C. Formal analysis: Gaona-López C. Funding acquisition: Gaona-López C. Investigation: Gaona-López C. Methodology: Gaona-López C. Project administration: Gaona-López C. Resources: Gaona-López C. Software: Gaona-López C. Supervision: Gaona-López C. Validation: Gaona-López C. Visualization: Gaona-López C. Writing – original draft: Gaona-López C. Writing – review & editing: Gaona-López C.
-
Conflict of interest
The author has no conflicts of interest to declare.
-
Acknowledgments
Carlos Gaona-López gratefully acknowledges the postdoctoral fellowship awarded by the Dirección General de Asuntos del Personal Académico (DGAPA), Universidad Nacional Autónoma de México (UNAM), during the period 2018–2020. He also expresses his sincere appreciation to the Departamento de Educación Continua of Instituto Nacional de Medicina Genómica (INMEGEN) for granting tuition-free scholarships that enabled his participation in several specialized training courses.
Supplementary information
Fig. 1.Chemical compounds related to treating Giardiasis and the potential inhibition of Giardia lamblia TATA-binding protein. (A) Chemical structures of the primary drugs currently used to treat Giardiasis: metronidazole, tinidazole, albendazole, mebendazole, and nitazoxanide. (B) Chemical structures of piceatannol and apigenin, 2 compounds identified in a previous study by our group as potential inhibitors of the TATA-binding protein of G. lamblia, based on molecular modeling tools.
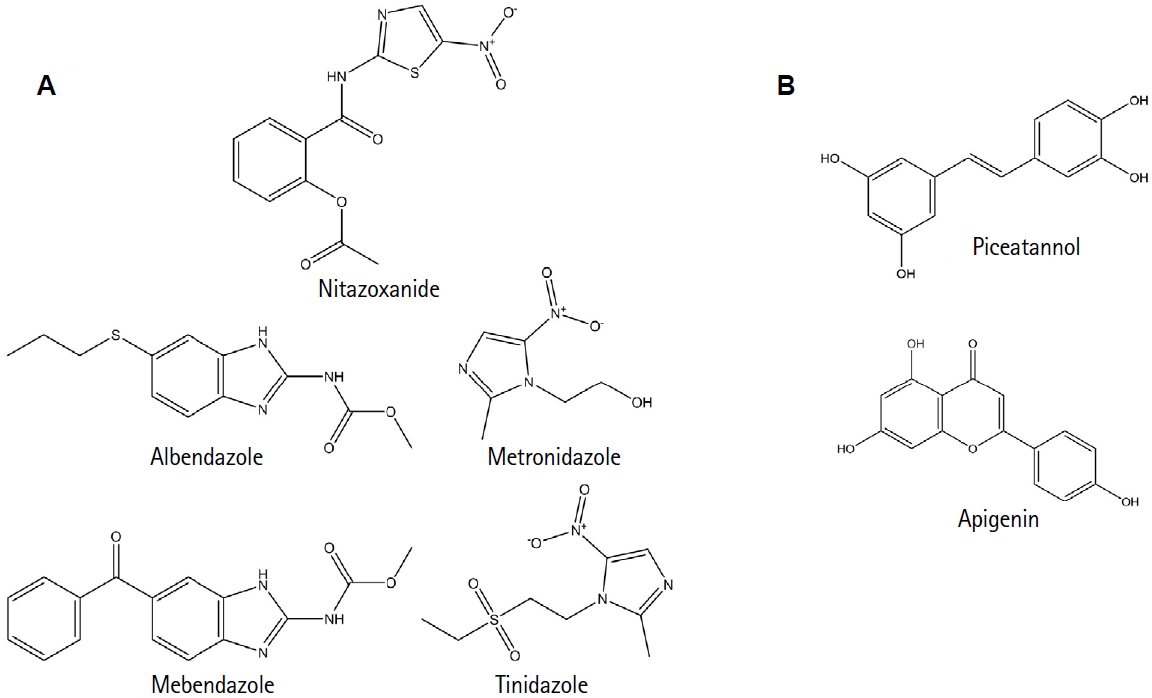
Fig. 2.Ultrastructural analysis of the nucleolus in Giardia lamblia trophozoites under control and actinomycin D-treated conditions. (A) Transmission electron microscopy image of untreated G. lamblia trophozoites showing intact nucleolar structure within the 2 nuclei; blue arrows indicate the nucleolus. (B) Trophozoites treated with actinomycin D (0.2 μg/ml) exhibit disorganization and dispersion of nucleolar material. This nucleolar disaggregation indicates nucleolar stress and reflects the transcriptional arrest of ribosomal RNA synthesis. The red box indicates the region where the nucleolus would usually be located. These structural changes provide additional evidence supporting the presence of a functional nucleolus in G. lamblia.
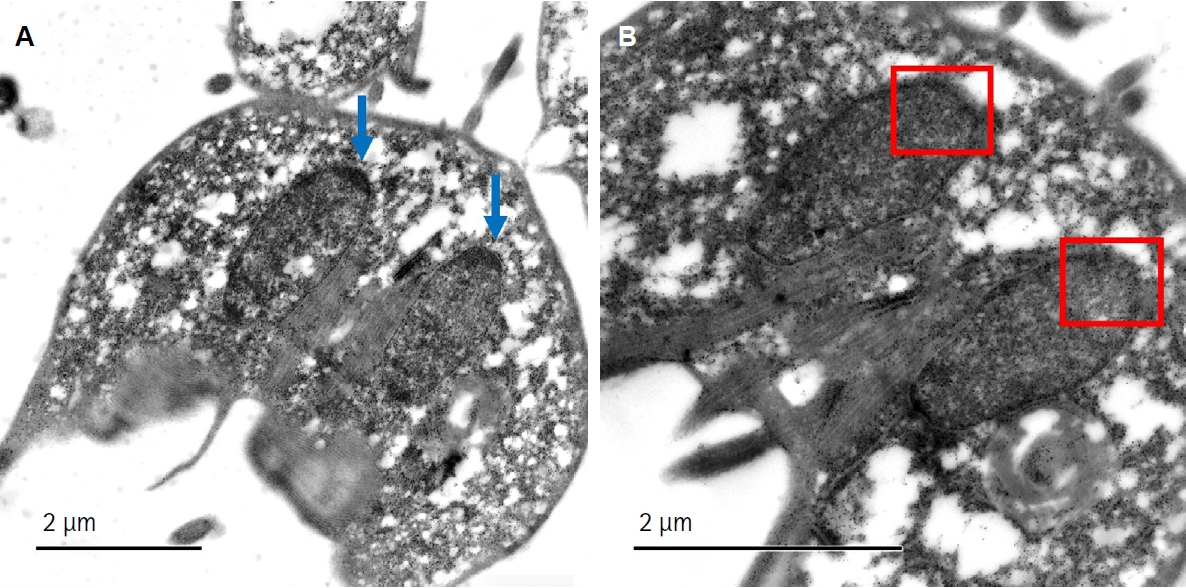

Fig. 3.Structural comparison between HsTBP and GlTBP. Surface representations of HsTBP (red) and GlTBP (green) are shown individually (top) and as a structural alignment (bottom). The human TATA-binding protein (TBP) structure corresponds to the crystallographic model PDB ID: 1JFI, whereas the 3D structure of Giardia lamblia TBP was obtained using AlphaFold 3. Both models were visualized and aligned using UCSF ChimeraX. The comparison highlights pronounced differences in surface topology and molecular volume between the 2 proteins. These structural deviations illustrate the high evolutionary divergence of G. lamblia TBP relative to its human counterpart, supporting its consideration as a potential selective drug target.
References
- 1. Cernikova L, Faso C, Hehl AB. Five facts about Giardia lamblia. PLoS Pathog 2018;14:e1007250. https://doi.org/10.1371/journal.ppat.1007250
- 2. Lanata CF, Fischer-Walker CL, Olascoaga AC, et al. Global causes of diarrheal disease mortality in children <5 years of age: a systematic review. PLoS One 2013;8:e72788. https://doi.org/10.1371/journal.pone.0072788
- 3. Savioli L, Smith H, Thompson A. Giardia and Cryptosporidium join the 'Neglected Diseases Initiative.'. Trends Parasitol 2006;22:203-8. https://doi.org/10.1016/j.pt.2006.02.015
- 4. Gaona-López C, Martínez-Vázquez AV, Villalobos-Rocha JC, Juárez-Rendón KJ, Rivera G. Analysis of Giardia lamblia nucleolus as drug target: a review. Pharmaceuticals (Basel) 2023;16:1168. https://doi.org/10.3390/ph16081168
- 5. Kotloff KL, Nataro JP, Blackwelder WC, et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case-control study. Lancet 2013;382:209-22. https://doi.org/10.1016/S0140-6736(13)60844-2
- 6. Argüello-García R, Leitsch D, Skinner-Adams T, Ortega-Pierres MG. Drug resistance in Giardia: mechanisms and alternative treatments for Giardiasis. Adv Parasitol 2020;17:201-82. https://doi.org/10.1016/bs.apar.2019.11.003
- 7. Escobedo AA, Cimerman S. Giardiasis: a pharmacotherapy review. Expert Opin Pharmacother 2007;8:1885-902. https://doi.org/10.1517/14656566.8.12.1885
- 8. Gaona-López C, Méndez-Álvarez D, Moreno-Rodríguez A, et al. TATA-binding protein-based virtual screening of FDA drugs identified new anti-Giardiasis agents. Int J Mol Sci 2024;25:6238. https://doi.org/10.3390/ijms25116238
- 9. Bendesky A, Menéndez D, Ostrosky-Wegman P. Is metronidazole carcinogenic? Mutat Res 2002;511:133-44. https://doi.org/10.1016/s1383-5742(02)00007-8
- 10. Dingsdag SA, Hunter N. Metronidazole: an update on metabolism, structure-cytotoxicity and resistance mechanisms. J Antimicrob Chemother 2018;73:265-79. https://doi.org/10.1093/jac/dkx351
- 11. IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. Ingested nitrate and nitrite, and cyanobacterial peptide toxins. International Agency for Research on Cancer; 2010.
- 12. Reiner DS, McCaffery M, Gillin FD. Sorting of cyst wall proteins to a regulated secretory pathway during differentiation of the primitive eukaryote, Giardia lamblia. Eur J Cell Biol 1990;53:142-53.
- 13. Marti M, Regös A, Li Y, et al. An ancestral secretory apparatus in the protozoan parasite Giardia intestinalis. J Biol Chem 2003;278:24837-48. https://doi.org/10.1074/jbc.M302082200
- 14. Adam RD. Giardia duodenalis: biology and pathogenesis. Clin Microbiol Rev 2021;34:e0002419. https://doi.org/10.1128/CMR.00024-19
- 15. Morrison HG, McArthur AG, Gillin FD, et al. Genomic minimalism in the early diverging intestinal parasite Giardia lamblia. Science 2007;317:1921-6. https://doi.org/10.1126/science.1143837
- 16. Narcisi EM, Glover CV, Fechheimer M. Fibrillarin, a conserved pre-ribosomal RNA processing protein of Giardia. J Eukaryot Microbiol 1998;45:105-11. https://doi.org/10.1111/j.1550-7408.1998.tb05077.x
- 17. Guo J, Chen Y, Zhou K, Li J. Distribution of rDNA in the nucleus of Giardia lamblia: detection by Ag-I silver stain. Biotech Histochem 2005;80:31-4. https://doi.org/10.1080/10520290500050981
- 18. Xin DD, Wen JF, He D, Lu SQ. Identification of a Giardia krr1 homolog gene and the secondarily anucleolate condition of Giardia lamblia. Mol Biol Evol 2005;22:391-4. https://doi.org/10.1093/molbev/msi052
- 19. Saraiya AA, Wang CC. snoRNA, a novel precursor of microRNA in Giardia lamblia. PLoS Pathog 2008;4:e1000224. https://doi.org/10.1371/journal.ppat.1000224
- 20. Jiménez-García LF, Zavala G, Chávez-Munguía B, et al. Identification of nucleoli in the early branching protist Giardia duodenalis. Int J Parasitol 2008;38:1297-304. https://doi.org/10.1016/j.ijpara.2008.04.012
- 21. Tian XF, Yang ZH, Shen H, Adam RD, Lu SQ. Identification of the nucleoli of Giardia lamblia with TEM and CFM. Parasitol Res 2010;106:789-93. https://doi.org/10.1007/s00436-009-1715-3
- 22. Hernandez-Verdun D. Assembly and disassembly of the nucleolus during the cell cycle. Nucleus 2011;2:189-94. https://doi.org/10.4161/nucl.2.3.16246
- 23. Best AA, Morrison HG, McArthur AG, Sogin ML, Olsen GJ. Evolution of eukaryotic transcription: insights from the genome of Giardia lamblia. Genome Res 2004;14:1537-47. https://doi.org/10.1101/gr.2256604
- 24. Sobell HM. Actinomycin and DNA transcription. Proc Natl Acad Sci U S A 1985;82:5328-31. https://doi.org/10.1073/pnas.82.16.5328
- 25. Sogin ML, Gunderson JH, Elwood HJ, Alonso RA, Peattie DA. Phylogenetic meaning of the kingdom concept: an unusual ribosomal RNA from Giardia lamblia. Science 1989;243:75-7. https://doi.org/10.1126/science.2911720
- 26. Hernandez N. TBP, a universal eukaryotic transcription factor? Genes Dev 1993;7:1291-308. https://doi.org/10.1101/gad.7.7b.1291
- 27. Parra-Marín O, López-Pacheco K, Hernández R, López-Villaseñor I. The highly diverse TATA box-binding proteins among protists: a review. Mol Biochem Parasitol 2020;239:111312. https://doi.org/10.1016/j.molbiopara.2020.111312
- 28. Santiago Á, Razo‐Hernández RS, Pastor N. The TATA‐binding protein DNA‐binding domain of eukaryotic parasites is a potentially druggable target. Chem Biol Drug Des 2020;95:130-49. https://doi.org/10.1111/cbdd.13630
Citations
Citations to this article as recorded by