Abstract
Malaria, caused by Plasmodium species, remains a major global health burden. The spleen is the central organ for clearance of infected red blood cells and regulation of immunity, yet paradoxically also serves as a site of parasite sequestration. Splenic fibroblasts may contribute to this process through adhesion mechanisms, but their role remains poorly defined. This study investigated P. yoelii tryptophan-rich antigen 7 (PyTRAg7), a member of the TRAg protein family, in splenic parasite–host interactions. TRAgs constitute a conserved protein family present in multiple Plasmodium species, including human malaria parasites, suggesting the relevance of PyTRAg7-associated mechanisms to human infection. Using protein binding assays and gene-edited parasites, PyTRAg7 was shown to interact with vimentin on mouse splenic fibroblasts, activating NF-κB p65 signaling and increasing ICAM-1 and integrin β1 expression. Deletion of PyTRAg7 reduced infected red blood cells adhesion to mouse splenic fibroblasts, lowered splenic parasite burden, delayed parasitemia onset, and prolonged host survival. Histological analysis showed preserved splenic architecture and reduced hemozoin deposition in the absence of PyTRAg7. The murine P. yoelii model was used due to its genetic tractability and its suitability for dissecting spleen-dependent sequestration mechanisms not easily studied in human malaria parasites. These findings identify PyTRAg7 as a key mediator of fibroblast–parasite interactions that promote cytoadherence and splenic remodeling, offering a potential target for malaria intervention.
-
Key words: Malaria, Plasmodium yoelii, fibroblasts, spleen, vimentin, NF-κB
Introduction
Malaria is a mosquito-borne infectious disease caused by
Plasmodium parasites and remains one of the leading challenges to global public health [
1]. During infection, the cytoadherence of infected red blood cells (iRBCs) contributes critically to malaria pathogenesis by causing microvascular obstruction, metabolic dysregulation, and the release of pro-inflammatory mediators [
2,
3]. A well-documented mechanism is the cytoadherence of iRBCs to vascular endothelial cells, which promotes sequestration in deep tissues and allows the parasites to evade splenic filtration [
4,
5]. Beyond endothelial cells, accumulating evidence indicates that iRBCs may also interact with diverse non-endothelial cell types across multiple organs, thereby facilitating immune evasion and persistent infection. In particular, splenic fibroblasts (SFs) are emerging as important contributors to parasite retention and remodeling of the splenic microenvironment [
6,
7].
The spleen plays a central role in malaria pathogenesis as both a site of iRBCs clearance and an immunological hub that orchestrates innate and adaptive immune responses [
8]. However, it is increasingly recognized that malarial parasites can exploit the splenic microenvironment as a niche for immune evasion [
9]. Recent studies suggest that SFs, traditionally viewed as structural components of the red pulp and marginal zone, may actively mediate adhesion, reorganize extracellular architecture, and form a physical “blood-spleen barrier” that facilitates parasite survival [
10,
11]. Yet, the molecular pathways that regulate fibroblast–parasite interactions remain poorly defined.
Parasite-exported proteins are key effectors of host-pathogen interactions [
12]. In
P. falciparum, PfEMP1 mediates adhesion to a range of endothelial receptors, including ICAM-1 and CD36, thereby facilitating vascular sequestration in a tissue-specific manner [
13]. In
P. vivax, proteins such as VIR14 and PvSRA interact with human SFs via ICAM-1 and ITGB1, respectively, thereby altering splenic structure and immune function [
11,
14]. In addition, tryptophan-rich antigens (TRAgs), such as PvTRAg23, have been shown to engage SFs and modulate extracellular matrix components like collagen I, further implicating these proteins in splenic remodeling and parasite persistence [
6]. Importantly, TRAgs are present not only in rodent malaria parasites but also in human-infecting species [
15], and several PvTRAgs exhibit strong immunogenicity and adhesive interactions with host cells [
16,
17], suggesting that TRAg-mediated mechanisms may be conserved contributors to malaria pathophysiology.
The TRAg protein family was first described in the rodent malaria parasite
P. yoelii [
18]. Rodent malaria models, particularly
P. yoelii, are widely used due to substantial similarities between murine and human immunobiology [
19]. Its lethal (17XL, YM) and non-lethal (17XNL, 17X) lines reproduce key disease phenotypes seen in
P. falciparum and
P. vivax—including high parasitemia, severe anemia, splenomegaly, and self-resolving infections [
20]. Combined with its genetic tractability, these features make
P. yoelii a powerful in vivo system for investigating spleen-dependent host–parasite interactions that cannot be readily examined in human malaria.
Among these proteins, PyTRAg7 is a surface-exported antigen. Its homolog, SLTRiP, has been linked to cellular immunity and adhesion, underscoring the broader functional significance of this protein family [
21]. Notably, several splenic-dependent PvTRAgs have been identified in
P. vivax, including PvTRAg5, PvTRAg16, PvTRAg23, PvTRAg30, and PvTRAg32, which are capable of interacting with SFs and modulating splenic functions [
7]. Despite these insights, whether PyTRAg7 contributes to immune evasion through interactions with SFs remains unknown.
Here, PyTRAg7 is identified as a novel mediator of host–parasite interactions in the spleen. It specifically binds to mouse SFs (MSFs) via surface vimentin, activates NF-κB p65 signaling, and upregulates adhesion molecules ICAM-1 and ITGB1. Functional deletion of PyTRAg7 reduces iRBCs adhesion, decreases splenic parasite burden, and attenuates virulence. These findings reveal a novel mechanism by which P. yoelii remodels the splenic microenvironment to promote immune evasion and disease progression.
Methods
Ethics statement
All procedures involving animals followed institutional and national guidelines for the care and use of laboratory animals and received prior approval from the Animal Ethics Committee of Jiangnan University (approval No. JN. No. 20200530b0301031). Female BALB/c mice (6 weeks old, 18–22 g) were maintained under specific-pathogen-free conditions with unrestricted access to food and water. Animals were randomly assigned to experimental groups and infected by intraperitoneal injection with 1×105 P. yoelii 17XL–iRBCs in 200 μl sterile saline. Parasitemia was monitored by Giemsa-stained blood smears. Mice were euthanized precisely on day 4 post-infection using CO₂ inhalation (gradual fill, ~20%–30% chamber volume/min) followed by cervical dislocation.
Cell culture
MSFs (Suzhou Bena Chuanglian Biotechnology) were grown in ScienCell Fibroblast Medium supplemented with 2% fetal bovine serum, 1% fibroblast growth supplement, and 1% penicillin–streptomycin. Cultures were maintained at 37°C in a humidified incubator with 5% CO₂. To enhance attachment, cultureware was pre-coated with poly-L-lysine (ScienCell) and incubated at 37°C overnight, rinsed twice with sterile water, and equilibrated with complete medium before seeding. Cells were subcultured when they reached ~95% confluence.
Recombinant protein production and purification
The coding sequences of pytrag7 (PlasmoDB accession PYYM_0701200) and pytrag1 (PlasmoDB accession PYYM_0625300) were codon-optimized for Escherichia coli and synthesized commercially (Tianlin Bio). Each optimized fragment was cloned into pET-30a to express PyTRAg7 or PyTRAg1 fused to an N-terminal thioredoxin and a C-terminal His tag to facilitate purification and detection. Both proteins were expressed in E. coli BL21 (DE3) pLysS (TransGen Biotech) under identical culture and induction conditions, and purified under non-denaturing conditions on Ni-Sepharose by YouLong Bio.
Western blotting
MSFs (5×106) were cultured to ~80% confluence and treated with or without PyTRAg7 for 48 h. Cells were lysed on ice for 30 min in RIPA buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1.0% NP-40 [IGEPAL CA-630], 0.5% sodium deoxycholate, 0.1% SDS, 1 mM EDTA) supplemented with protease and phosphatase inhibitor cocktails (Beyotime). Lysates were clarified at 12,000× g for 10 min at 4°C, denatured by boiling, resolved on 10.0% SDS-PAGE, and transferred to PVDF membranes. Membranes were blocked in TBST containing 5% skim milk (2 h, room temperature [RT]) and incubated overnight at 4°C with the following primary antibodies: anti-His (1:1,000, Abcam), anti-GFP (1:1,000, Abcam), NF-κB p65 (1:1000, CST), phospho-NF-κB p65 (1:1,000, CST), ERK (1:1,000, CST), phospho-ERK (1:1,000, CST), and GAPDH (1:5,000, Abcam). After TBST washes, HRP-conjugated secondary antibodies (1:5,000, Abcam) were applied for 1 h at RT. Signals were developed with an ECL kit (NCM) and quantified using ImageJ (NIH).
RNA extraction and quantitative real-time PCR
Total RNA was extracted from MSFs (5×10
6) or pulverized spleen tissue using the manufacturer’s protocol (ES Science). Genomic DNA was removed with gDNA Remover Mix, and cDNA was synthesized using 5× HiFiScript RT MasterMix (CWBIO). qPCR reactions (triplicates per sample) were set up with 2× TB Green Premix Ex Taq II (Takara) and run on a LightCycler 480 II (Roche). Cycling: 95°C for 30 sec; 40 cycles of 95°C for 5 sec and 60°C for 30 sec; followed by 95°C for 5 sec, 60°C for 1 min, and 50°C for 30 sec. GAPDH served as the internal reference. Primer sequences are listed in
Supplementary Table S1.
MSFs (5×106) were incubated with rabbit anti-His antibody (1:300, Abcam) for 30 min at 4°C, washed twice with phosphate-buffered saline (PBS), and stained with Alexa Fluor 647–conjugated goat anti-rabbit IgG (1:2,000, Abcam) for 1 h at 4°C protected from light. Samples were analyzed on a FACScan flow cytometer (Becton Dickinson). The proportions of His-positive or vimentin-positive cells were determined using FlowJo (FlowJo LLC).
HEK293T culture and transient transfection
HEK293T cells were maintained in DMEM containing 10% fetal bovine serum (Gemini Bio-Products), 100 U/ml penicillin, and 100 μg/ml streptomycin at 37°C with 5% CO₂. For immunoprecipitation, cells seeded in 6-cm dishes were transfected with 5 μg each of pEGFP-C1-vimentin, pEGFP-C1-RACK1, pEGFP-C1-Phb2, or pEGFP-C1-vector using Lipofectamine (Yeasen) according to the manufacturer’s instructions. After 24 h, transfection efficiency was examined by fluorescence microscopy (Nikon). Cells were lysed with NP-40 buffer (Beyotime), and expression of vimentin, RACK1, and Phb2 was confirmed by Western blot.
Pulldown assay
For screening candidate interactors of PyTRAg7, 200 μg His-tagged PyTRAg7 was bound to 500 μl Ni-NTA agarose (Qiagen) pre-equilibrated in buffer (50 mM NaH₂PO₄, 300 mM NaCl, 20 mM imidazole, pH 8.0). After overnight incubation at 4°C on a rotator, the resin was washed with equilibration buffer and incubated overnight with 200 μg MSFs membrane proteins extracted using the Mem-PER Plus kit (Thermo Fisher Scientific). Beads were washed 3 times with pre-chilled isotonic buffer followed by hypertonic buffer to remove non-specific binders, and bound proteins were eluted stepwise with imidazole at 100, 200, 300, 400, and 500 mM (same buffer base as above; 500 μl per step, 15 min gentle rotation at 4°C, then 12,000× g, 5 min, 4°C). Eluates were resolved by SDS-PAGE and visualized by silver staining (Fast Silver Stain Kit, Beyotime). Specific bands were excised and submitted to Shanghai Applied Protein Technology Company for liquid chromatography-mass spectrometry analysis.
To assess PyTRAg7–vimentin interaction, 200 μg PyTRAg7 was immobilized on 500 μl Ni-NTA agarose for 3 h at 4°C, washed, and incubated overnight with lysates from HEK293T cells expressing pEGFP-C1-vimentin. After extensive washing with 20 bed volumes of equilibration buffer, bound proteins were eluted; 100 μl eluate was mixed with 25 μl 5× SDS reducing buffer, boiled for 8 min, and analyzed by Western blot.
Immunofluorescence colocalization
MSFs were seeded onto 15-mm confocal culture dishes and grown to ~70%–80% confluence. At ~80% density, cells were incubated in a 37°C incubator with His-tagged recombinant protein (PyTRAg7-His or PyTRAg1-His; 10 μg/ml) for 2 h; the BC group received vehicle only. Cells were rinsed 3× with pre-chilled PBS and fixed in 4% paraformaldehyde for 30 min at RT, followed by PBS washes (3×). After blocking in 10% bovine serum albumin (BSA) for 30 min at RT, primary antibodies diluted in 3% BSA were applied overnight at 4°C: mouse anti-His (1:100, Abcam) to detect the recombinant protein and rabbit anti-vimentin (1:100, Zenbio) for surface vimentin. Following PBS washes (3×), species-matched secondary antibodies diluted in 3% BSA were applied for 1 h at RT in the dark: Alexa Fluor 488 anti-rabbit (1:1,000) and Alexa Fluor 647 anti-mouse (1:1,000). Nuclei were counterstained with DAPI (1:1,000 in PBS, 10 min, RT, dark) and washed 5× with PBS. Dishes were kept in PBS for imaging. Images were acquired on a confocal microscope under identical laser and detector settings across all groups, using sequential single-channel acquisition (488 → 647) to avoid bleed-through.
Generation of ΔTRAg7 parasites
A
pytrag7 knockout strain (ΔTRAg7) was generated using a CRISPR-Cas9-based double-crossover homologous recombination strategy, as established in a parallel study by our group (unpublished data). Briefly, ~1 kb fragments corresponding to the 5′ UTR and 3′ ORF of
pytrag7 were amplified from
P. yoelii 17XL genomic DNA and inserted into the pPyCrispr vector using standard restriction enzyme cloning. The resulting replacement cassette was released and transfected into schizont-stage parasites via Amaxa nucleofection. Transgenic parasites were selected in mice with daily pyrimethamine administration (1 mg/kg), and drug-resistant clones were obtained by serial dilution. Two independent ΔTRAg7 clones were isolated and exhibited identical growth and molecular profiles. One representative clone was selected for phenotypic analysis in this study. Gene disruption was confirmed by diagnostic PCR and RT-PCR using primer combinations listed in
Supplementary Tables S2 and
S3.
Peripheral blood was collected by intracardiac puncture from 17XL-infected mice at day 4 post-infection into EDTA/PBS. Pelleted cells were washed 3 times with RPMI-based adhesion medium (RPMI 1640 with L-glutamine, 20 mM HEPES, 50 μg/ml gentamicin; pH 6.8). Mature parasite-infected RBCs were enriched on a 55% Histodenz/PBS gradient. For adhesion, duplicate wells received 1×106 iRBCs in 300 μl adhesion medium and were incubated at 37°C for 1 h. Slides were washed by 10 serial immersions in adhesion medium, air-dried, methanol-fixed, and Giemsa-stained. Counts were averaged over 150 high-power fields (100× objective) per well.
Quantification of adherent iRBCs
For each condition (normal RBCs, wild-type [WT] iRBCs, ΔTRAg7 iRBCs), ≥3 biological replicates were analyzed. Per replicate, 3 slides were prepared and 5 random fields per slide were imaged at 1,000× oil immersion under identical settings (scale bar indicated). An adherent iRBC was defined as an iRBC in direct contact with the MSF membrane and remaining after standardized washes; touching but non-adherent cells and out-of-focus events were excluded. Using ImageJ, the number of adherent iRBCs per individual MSF was recorded in a blinded manner until ≥150 MSFs were counted per replicate. The outcome metric was the mean number of adherent iRBCs per MSF for each replicate.
Hematoxylin-eosin staining
Spleen sections (20 μm) were stained using the HE Staining Kit (Solarbio, G1120) according to the kit protocol. Briefly, sections were stained with hematoxylin for 1 min, differentiated in acid alcohol for 30 s, counterstained with eosin for 50 s, dehydrated through 95% and 100% ethanol, cleared in xylene, and mounted. Images were acquired on a Nikon Ci-L microscope or an Olympus SLIDEVIEW VS200 (Olympus) and analyzed with ImageJ.
Statistical analysis
For normally distributed data with more than 2 groups, one-way ANOVA was applied; pairwise comparisons were conducted using Student’s t-test. Analyses were performed in GraphPad Prism version 8 (GraphPad Software). Differences were considered statistically significant at P<0.05 (*P<0.05, **P<0.01, ***P<0.001).
Results
Expression and purification of PyTRAg7
The full-length cDNA of the
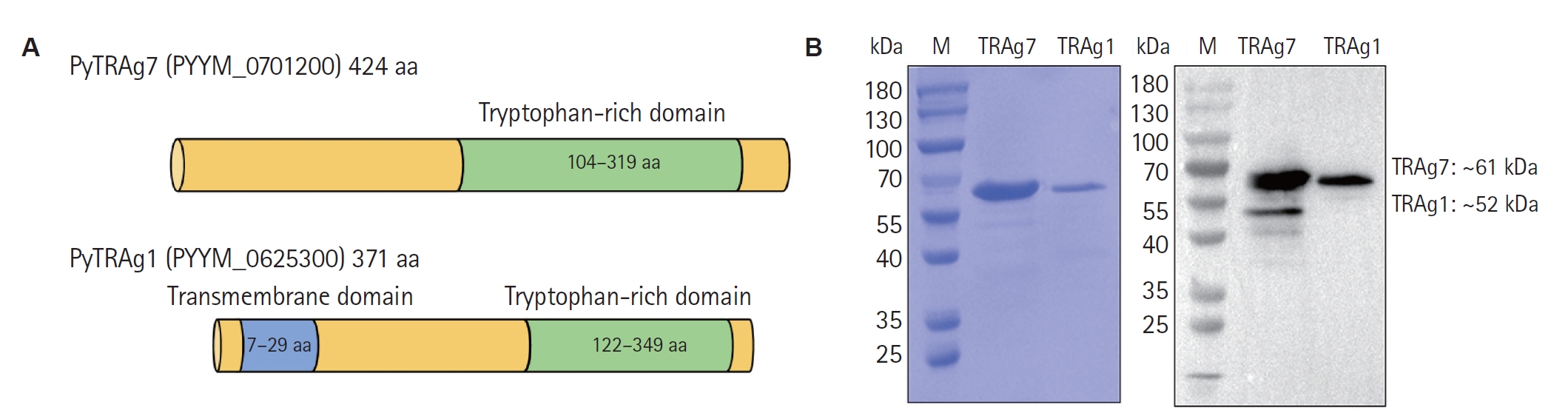
PYYM_0701200 gene encodes the PyTRAg7 protein comprising 424 amino acids, with a characteristic tryptophan-rich domain spanning residues 104–319. PyTRAg7 (full-length, aa 1–424) was cloned into pET-30a and expressed in
E. coli as a His-tagged protein (
Fig. 1A). In parallel, the homolog PyTRAg1 (371 aa) contains a signal peptide (aa 1–28) and a transmembrane region (aa 7–29) that are unlikely to be properly recognized and folded in the
E. coli system. Therefore, these 2 domains were removed during gene amplification. Both recombinant proteins were purified under identical conditions, subjected to endotoxin removal, and used for downstream assays. SDS-PAGE and Western blot analyses confirmed that both recombinant PyTRAg7 and PyTRAg1 were obtained as single distinct bands, with molecular weights consistent with their predicted sizes (
Fig. 1B).
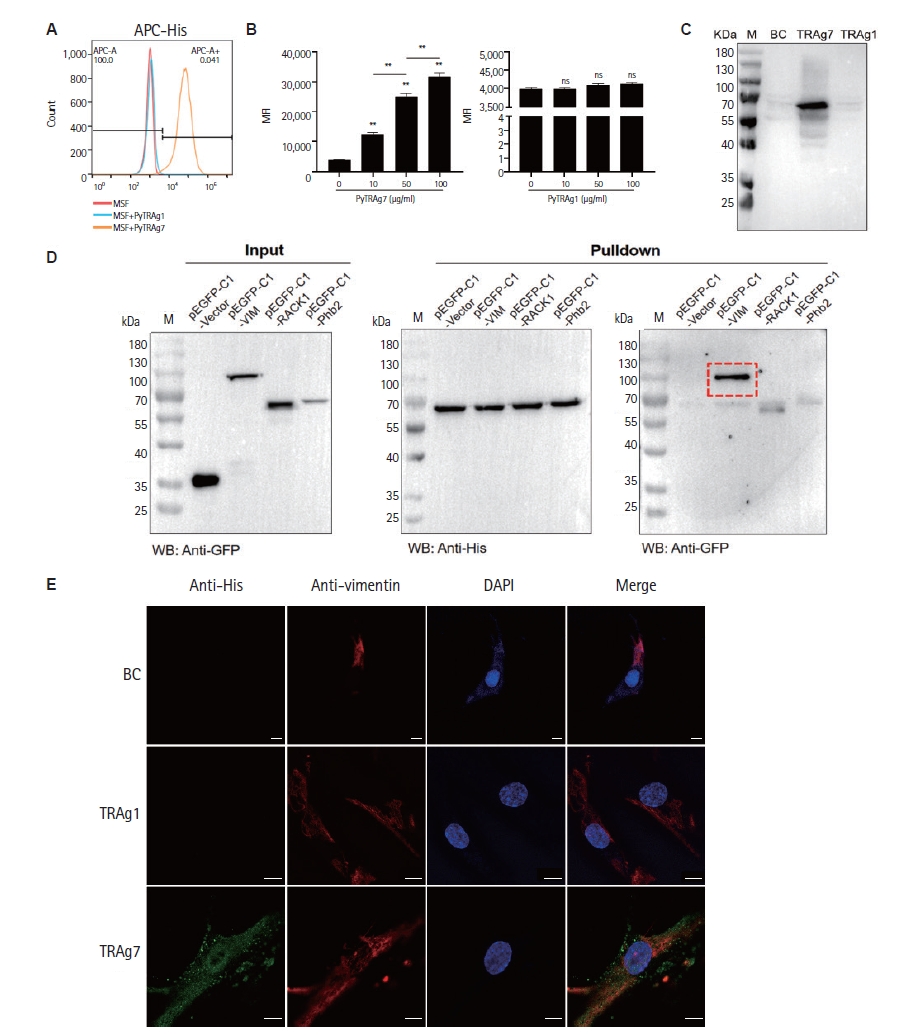
Flow cytometry showed a clear rightward shift of APC-His on MSFs with PyTRAg7 (100 μg/ml) but not with PyTRAg1 (100 μg/ml) (
Fig. 2A). Quantification across 0, 10, 50, 100 μg/ml revealed a dose-dependent increase in mean fluorescence intensity for PyTRAg7, whereas PyTRAg1 was unchanged (
Fig. 2B). In the Western blot–based binding assay (
Fig. 2C), 10 μg/ml recombinant protein produced a strong His signal for PyTRAg7 but not for PyTRAg1. Subsequently, differential bands were isolated using nickel column affinity chromatography and silver staining, followed by protein profiling to identify potential binding receptors. Among the candidate proteins identified, vimentin emerged with the highest score (
Table 1), suggesting its role as the likely binding receptor for PyTRAg7 in MSFs. Consequently, 293T cells were transfected with the pEGFP-C1-vimentin eukaryotic plasmid, and a His-pulldown assay was conducted post-verification of vimentin expression. As depicted in
Fig. 2D, a pronounced interaction between PyTRAg7 and vimentin was observed. To visualize this interaction on MSFs, immunofluorescence colocalization showed robust anti-His signal in the PyTRAg7 group that overlapped with vimentin filaments, whereas the BC and PyTRAg1 groups were negative under identical acquisition settings (
Fig. 2E).
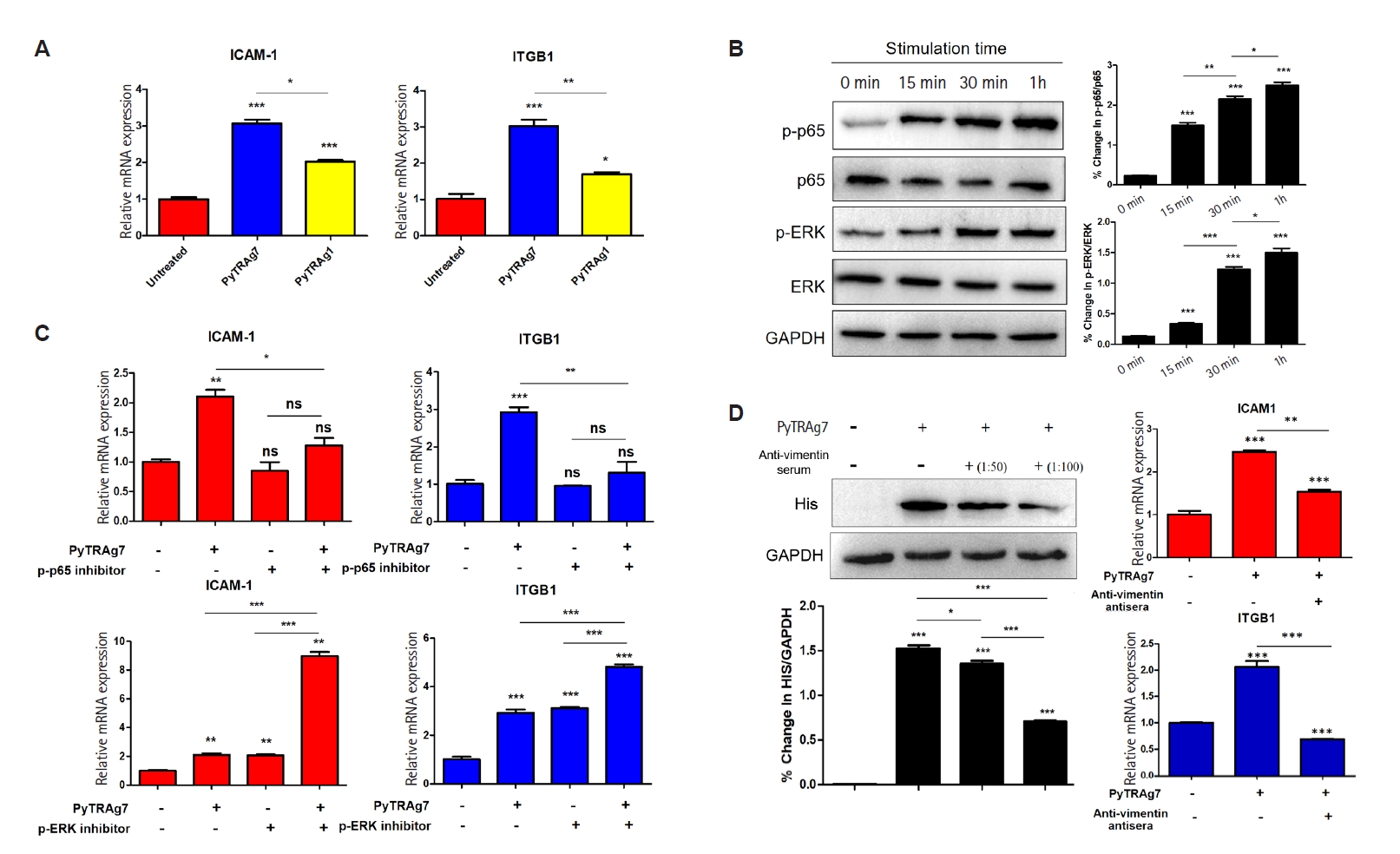
MSFs cultured with recombinant PyTRAg7 for 48 h displayed a marked increase in the mRNA levels of ICAM-1 and ITGB1, whereas PyTRAg1 induced only a modest elevation of these transcripts (
Fig. 3A). The weaker response to PyTRAg1 likely reflects a mild nonspecific effect of recombinant protein exposure, in contrast to the robust induction driven by PyTRAg7. Exposure of MSFs to PyTRAg7 led to progressive phosphorylation of both signaling proteins over time (
Fig. 3B), indicating their activation in response to PyTRAg7. When pathway-specific inhibitors were applied—Bay 11-7082 for NF-κB and SCH772984 for ERK—only NF-κB inhibition resulted in a marked suppression of ICAM-1 and ITGB1 expression (
Fig. 3C), pointing to NF-κB p65 as the key downstream effector. The functional involvement of vimentin was further examined using rabbit anti-vimentin serum to block receptor accessibility. Pre-incubation with the antibody (1:100 dilution) substantially reduced PyTRAg7 binding to MSFs (
Fig. 3D, left), accompanied by decreased phosphorylation of p65 and reduced mRNA levels of both adhesion molecules (
Fig. 3D, right). These findings support a model in which PyTRAg7 engages surface vimentin to activate the NF-κB p65 signaling cascade, ultimately enhancing expression of ICAM-1 and ITGB1 in MSFs.
A
pytrag7 knockout strain (ΔTRAg7) was generated using a CRISPR-Cas9-based double-crossover homologous recombination strategy, as previously established in a parallel study by our group. Genotyping PCR and Western blot confirmed complete disruption of the
pytrag7 locus and loss of PyTRAg7 protein expression (
Supplementary Fig. S1).
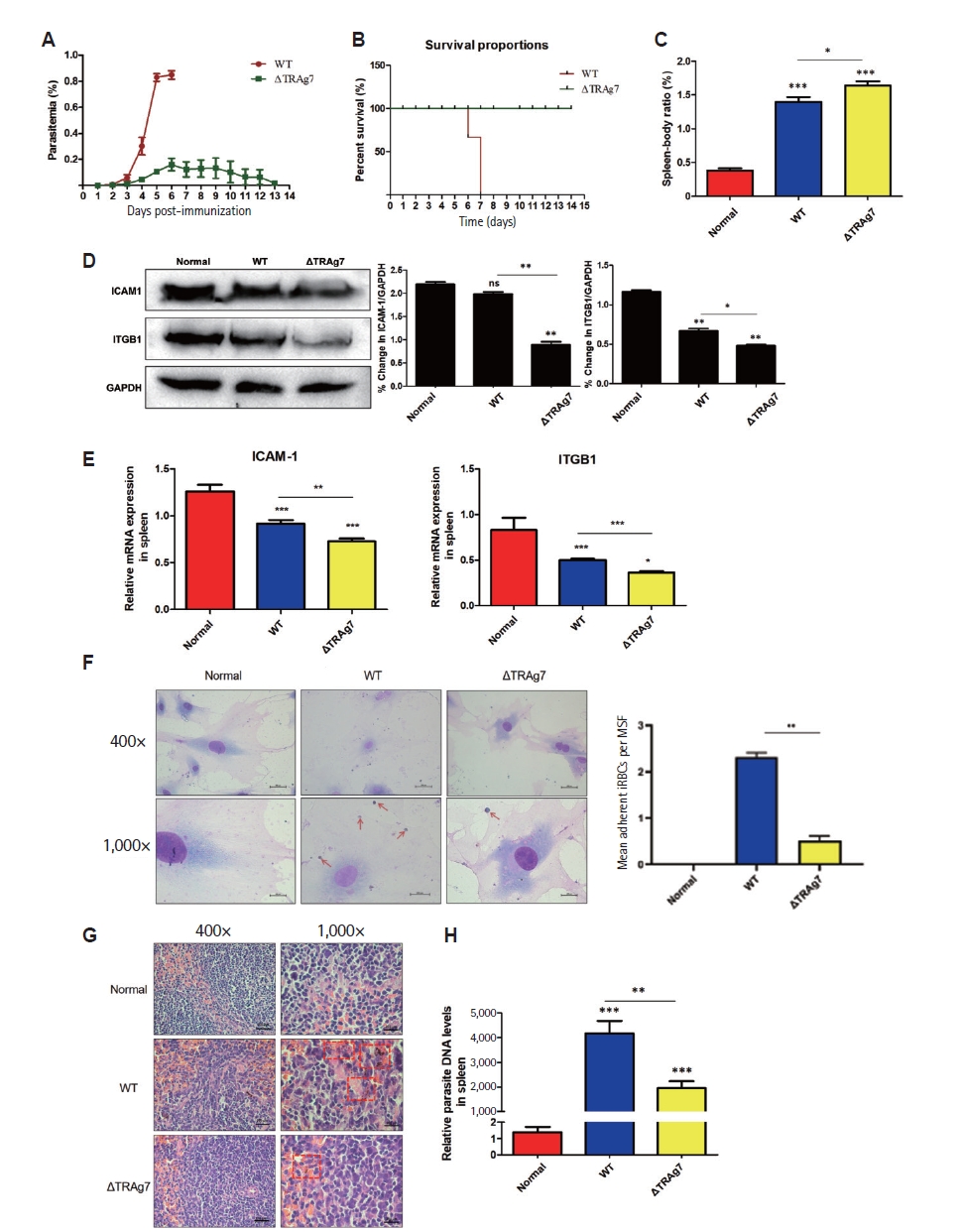
The impact of PyTRAg7 on parasite virulence was examined using BALB/c mice infected with either WT or ΔTRAg7
P. yoelii strains. Mice infected with WT parasites developed rapidly increasing parasitemia, reaching ~83% by day 6, followed by complete mortality on day 7 (
Fig. 4A,
B). In contrast, ΔTRAg7-infected mice showed delayed onset of parasitemia and gradual clearance, with no deaths observed over 14 days, highlighting the critical role of PyTRAg7 in driving acute infection. Given the spleen’s role in filtering iRBCs and regulating immune responses, spleen/body weight ratios were evaluated. Mice infected with ΔTRAg7 parasites displayed significantly higher splenic indices than both WT-infected and uninfected animals (
Fig. 4C).
The expression of host adhesion molecules was next assessed to clarify the potential mechanism underlying splenic differences. Both protein and mRNA levels of ICAM-1 and ITGB1 were markedly reduced in ΔTRAg7-infected spleens compared to WT counterparts (
Fig. 4D,
E), indicating that PyTRAg7 may enhance parasite retention by modulating host adhesion pathways. To directly evaluate adhesion capacity, an ex vivo assay was conducted using MSFs and merozoite-enriched iRBCs. Representative micrographs show abundant attachment of WT iRBCs to MSFs, whereas ΔTRAg7 iRBCs display visibly reduced binding and normal RBCs show negligible adhesion (
Fig. 4F, left). Quantification of adhesion (mean adherent iRBCs per MSF) confirmed a significant increase for WT compared with ΔTRAg7 and normal controls (
Fig. 4F, right). Histopathological analysis revealed clear architectural damage in WT-infected spleens, characterized by disrupted red and white pulp boundaries, heavy hemozoin deposition, and abundant parasite bodies. In contrast, ΔTRAg7-infected spleens maintained relatively intact structure and exhibited substantially less pigment accumulation (
Fig. 4G). Consistent with these findings, qPCR analysis of
Plasmodium-specific transcripts confirmed a markedly lower parasite burden in ΔTRAg7-infected spleens compared to WT controls at equivalent time points (
Fig. 4H). Together, these data demonstrate that PyTRAg7 facilitates
P. yoelii virulence by promoting iRBCs adhesion to SFs, thereby contributing to parasite sequestration and immune evasion within the spleen.
Discussion
Immune evasion represents a central strategy for
Plasmodium survival within the host [
22,
23]. Beyond serving as the primary site for clearance of iRBCs, the spleen also provides a niche that parasites exploit to avoid immune elimination [
9,
24]. Accumulating studies further suggest that
Plasmodium exported proteins can interact with host SFs and play an important role in this process [
7,
14]. Building on this concept, the present study identifies PyTRAg7 as a novel parasite effector that mediates immune evasion through direct engagement with SFs.
This study provides new evidence for the role of PyTRAg7 in splenic pathophysiology. PyTRAg7 was found to interact with vimentin on the surface of MSFs, triggering activation of NF-κB p65 signaling and subsequent upregulation of the adhesion molecules ICAM-1 and ITGB1. These molecular changes imply that PyTRAg7 functionally reprograms MSFs, enabling them to contribute actively to splenic microenvironment remodeling rather than serving only structural roles. These findings align with previous studies on TRAg family members. PvTRAg23 in
P. vivax engages vimentin on SFs, activating NF-κB signaling and driving pro-inflammatory responses while suppressing type I collagen secretion [
6]. Although downstream outcomes differ, such observations collectively suggest that TRAg proteins may broadly mediate parasite-host cell adhesion and microenvironmental modulation. The functional clustering of PyTRAg7 homologs, such as SLTRiP in
P. berghei, further supports a role for this family in adhesion-related processes [
21]. Proteins belonging to the same family are generally encoded by related genes and are often presumed to share functional similarities [
25,
26]. Together, these results highlight TRAgs as versatile mediators of host-parasite interactions beyond classical immune cell engagement.
At the mechanistic level, ICAM-1 and ITGB1 are critical adhesion molecules expressed in leukocytes, endothelial cells, and fibroblasts, and are increasingly recognized for their roles in malaria pathogenesis [
11,
14,
27,
28]. Their induction in MSFs by PyTRAg7 enhances iRBCs adhesion and may disrupt cell-matrix interactions within the splenic microenvironment, thereby impairing blood filtration and pathogen clearance. Vimentin, identified as the receptor for PyTRAg7, is a multifunctional cytoskeletal protein that often facilitates cellular invasion through ligand engagement [
29,
30]. Our localization analysis further demonstrates that PyTRAg7 colocalizes with surface vimentin on MSFs, providing direct spatial evidence for this receptor–ligand interaction. Its engagement by PyTRAg7 underscores a broader strategy by which pathogens exploit host cytoskeletal components to manipulate signaling pathways. Importantly, NF-κB–mediated responses induced by PyTRAg7 may exert dual effects: promoting parasite sequestration and immune evasion while also driving inflammatory pathology. This mechanism echoes prior findings that PySRA activates NF-κB signaling via CD68 in macrophages [
31], suggesting that diverse
Plasmodium exported antigens may converge on NF-κB signaling through distinct pathways to cooperatively shape the host microenvironment.
PyTRAg7 knockout parasites were generated to directly assess its functional role. ΔTRAg7 parasites exhibited reduced adhesion to MSFs, decreased splenic parasite loads, preserved splenic architecture, and significantly prolonged host survival. Histological analysis revealed diminished hemozoin deposition and parasite accumulation in the spleen, underscoring the contribution of PyTRAg7 to splenic sequestration and pathology. Importantly, although both WT and ΔTRAg7 infected mice developed splenomegaly, the spleens of ΔTRAg7 infected animals were noticeably larger. Rather than indicating exacerbated pathology, this enlargement is more consistent with compensatory splenic expansion driven by heightened immune activation. In the absence of PyTRAg7-mediated sequestration, parasites are less able to remain hidden within splenic niches and instead circulate through the spleen, where they are more effectively detected and cleared. Previous studies have shown that repeated or intensified exposure of the spleen to parasitized erythrocytes can promote expansion of the reticuloendothelial system and macrophage populations, a hallmark of malaria-associated splenomegaly [
32,
33]. Increased clearance demand can also induce extramedullary erythropoiesis and immune cell proliferation—processes known to contribute to spleen enlargement during malaria and other infectious conditions [
34]. Thus, the more pronounced splenomegaly observed in ΔTRAg7-infected mice likely reflects a more robust and effective splenic response to circulating parasites rather than more severe disease. Given the established role of the spleen as a reservoir for persistent
Plasmodium infection [
35], these results support a model in which PyTRAg7 reshapes the splenic microenvironment to facilitate long-term parasite persistence and transmission.
In summary, this study identifies PyTRAg7 as a parasite-derived effector that binds to vimentin on MSFs, activates NF-κB signaling, and induces the expression of ICAM-1 and ITGB1, thereby enhancing parasite adhesion and promoting splenic retention. These findings broaden the current understanding of
Plasmodium host interactions and underscore PyTRAg7 as a promising molecular target for therapeutic intervention. Although the molecular mechanism underlying PyTRAg7–MSF engagement is well supported by in vitro and in vivo evidence, additional studies will be required to fully establish the physiological relevance of these interactions in different stages of infection and across parasite strains. In particular, the extent to which TRAg-mediated fibroblast modulation is conserved in human malaria parasites remains an important question, and future work incorporating structural analysis of the PyTRAg7–vimentin interface, single-cell profiling of MSFs responses, and humanized spleen models may provide deeper mechanistic insight. Based on these data, a conceptual model is proposed to illustrate how PyTRAg7 mediates immune evasion within the spleen during
P. yoelii infection (
Fig. 5).
Notes
-
Author contributions
Conceptualization: Han S, Cheng Y. Data curation: Zhu Y, Zhang H, Sun Y. Formal analysis: Zhu Y, Zhang H, Sun Y. Funding acquisition: Cheng Y. Investigation: Zhu Y, Zhang H, Sun Y. Methodology: Zhu Y, Zhang H, Sun Y. Project administration: Han S, Cheng Y. Resources: Cheng Y. Software: Sun Y. Supervision: Sun Y, Han S, Cheng Y. Validation: Zhu Y, Zhang H, Sun Y. Visualization: Zhu Y, Zhang H. Writing – original draft: Zhu Y, Zhang H. Writing – review & editing: Sun Y, Han S, Cheng Y.
-
Conflict of interest
The authors have no conflicts of interest to declare.
-
Funding
This work was supported by the Bill & Melinda Gates Foundation (No. INV-061480), the National Natural Science Foundation of China (Grant No. 82560401 and 82502749), the China Postdoctoral Science Foundation (No. 2023M731343), the Natural Science Foundation of Jiangsu Province (No. BK20231494), the Scientific Research Program of Jiangsu Province Health Commission (No. x202321), the Top Talent Support Program for Young and Middle-aged People of Wuxi Health Committee (No. HB2023039), the Open Topic of the National Health Commission Key Laboratory of Parasite and Vector Biology (No. NHCKFKT2024-6), the Medical Discipline Development Fund of the School of Medicine, Jiangnan University (No. YXXK2024092604), and Wuxi Medical Key Discipline (No. ZDXK2021002).
-
Acknowledgments
The authors gratefully acknowledge the administrative support and technical assistance provided by laboratory members, as well as constructive discussions and critical reading of the manuscript by colleagues.
Supplementary information
Fig. 1.Expression and purification of recombinant PyTRAg7 and PyTRAg1. (A) Schematic of PyTRAg7 (424 aa) showing the tryptophan-rich region (aa 104–319). The full-length cDNA (aa 1–424) was cloned into pET-30a for expression in Escherichia coli. In parallel, PyTRAg1 (371 aa) was expressed as its soluble ectodomain (aa 30–371) after removal of the signal peptide (aa 1–28) and the predicted transmembrane segment (aa 7–29). (B) Purification and identity of recombinant proteins by SDS-PAGE (left) and anti-His Western blot (right). Recombinant PyTRAg7 (~61 kDa), and PyTRAg1 (~52 kDa). M, marker.
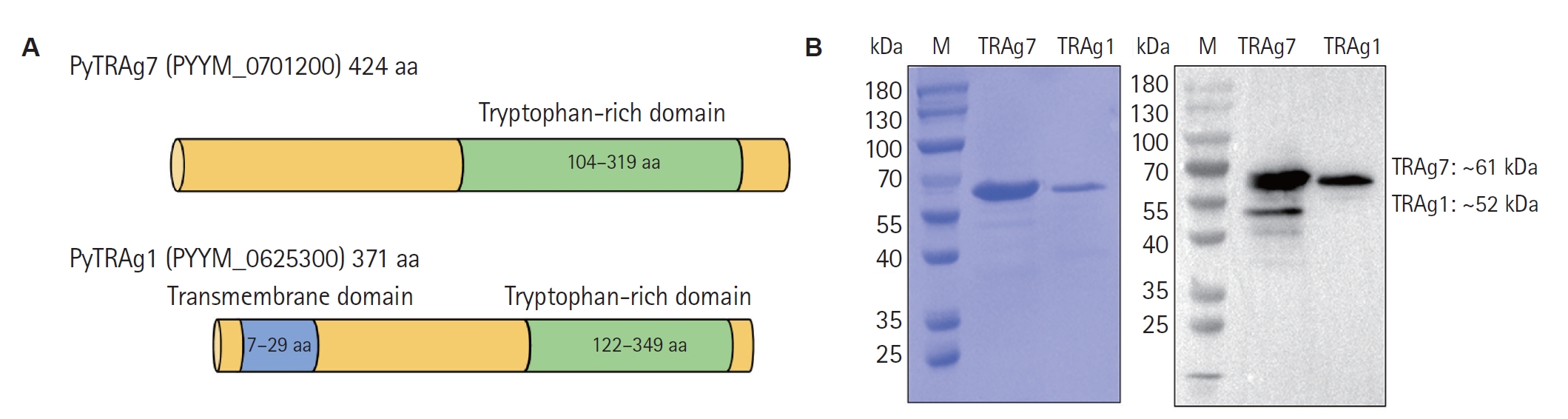
Fig. 2.Vimentin mediates PyTRAg7 binding on mouse splenic fibroblasts (MSFs). (A) Representative flow-cytometry histograms (APC–His) of MSFs incubated with PyTRAg7 or PyTRAg1 (100 μg/ml each) versus isotype control. The APC⁺ gate and %APC⁺ are indicated. (B) Quantification of APC–His signal as mean fluorescence intensity across doses (0, 10, 50, 100 μg/ml; n=3 independent experiments). PyTRAg7 shows a dose-dependent increase, whereas PyTRAg1 is unchanged. (C) Western blot–based binding assay: MSFs incubated with 10 μg/ml recombinant protein; a strong His signal is detected for PyTRAg7 but not PyTRAg1. (D) Binding receptor detection. 293T cells were transfected with GFP-tagged constructs, lysed, and the lysates were incubated with Ni-NTA–immobilized PyTRAg7 overnight. Eluates were probed with anti-His (to detect PyTRAg7) and anti-GFP (to detect vimentin), supporting vimentin as the PyTRAg7-binding receptor. (E) Immunofluorescence colocalization on MSFs. Cells were incubated with vehicle (BC), PyTRAg1-His or PyTRAg7-His (10 μg/ml, 2 h, 37°C), rinsed, fixed (4% PFA), and stained with anti-His (green) and anti-vimentin (red); nuclei were counterstained with DAPI (blue). Images were acquired under identical settings with sequential single-channel capture. PyTRAg7 shows clear overlap with vimentin, whereas BC and PyTRAg1 are negative. ns, not significant; M, marker; BC, bead control. Scale bars=10 μm.
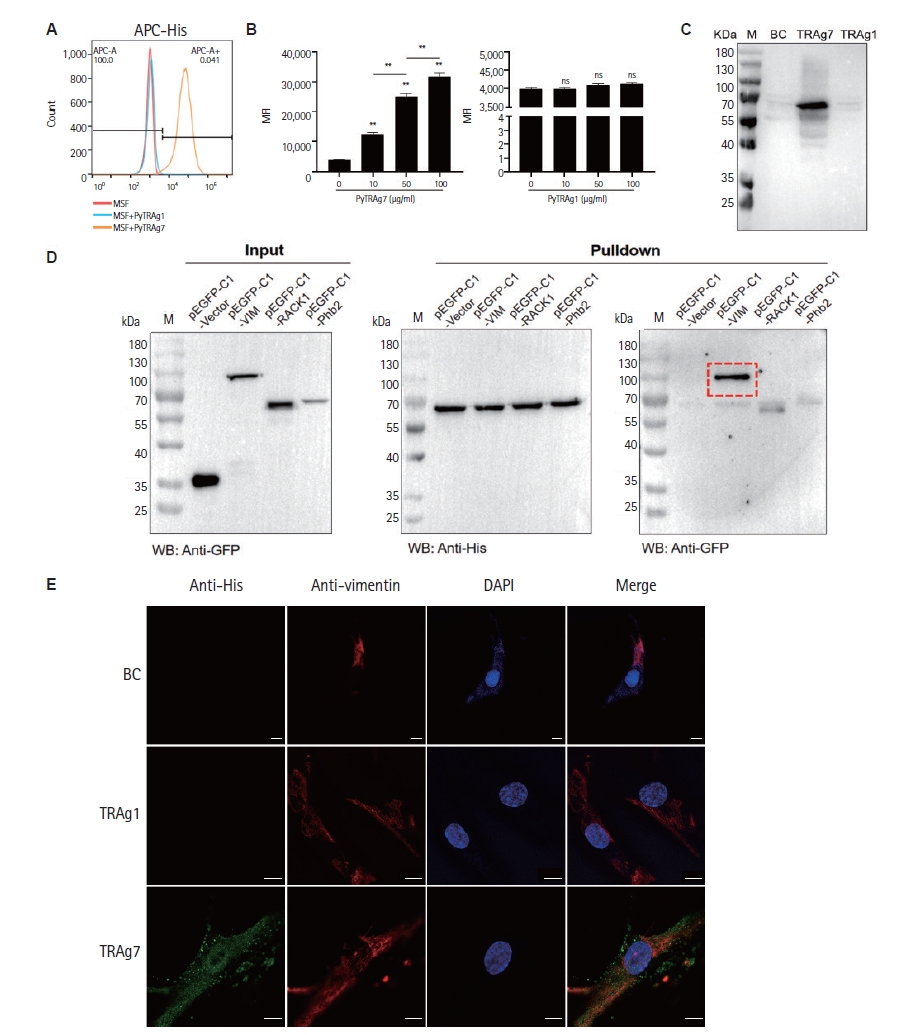
Fig. 3.PyTRAg7 mediates mouse splenic fibroblasts (MSFs) adhesion molecule production through activation of the NF-κB pathway. (A) Detection of transcription levels of MSFs adhesion molecules. MSFs treated with PyTRAg7 or PyTRAg1 were analyzed by qPCR for ICAM-1 and ITGB1 mRNA (n=3 independent experiments). (B) Signal path activation detection. Time-course immunoblots of NF-κB p65 and ERK in PyTRAg7-stimulated MSFs; GAPDH served as loading control. (C) Regulatory path detection. MSFs were pretreated with pathway inhibitors and then exposed to PyTRAg7; ICAM-1 and ITGB1 transcripts were quantified by qPCR. “+” indicates inhibitor/protein added; “–” indicates not added. (D) Detection of combined action. Both rabbit vimentin serum pretreated and untreated MSFs were stimulated with PyTRAg7, the mRNA expression of ICAM-1 and ITGB1 in each group was detected compared with the untreated group. Statistics: one-way ANOVA with Dunnett’s multiple comparisons test (nsP≥0.05; *P<0.05, **P<0.01, ***P<0.001).
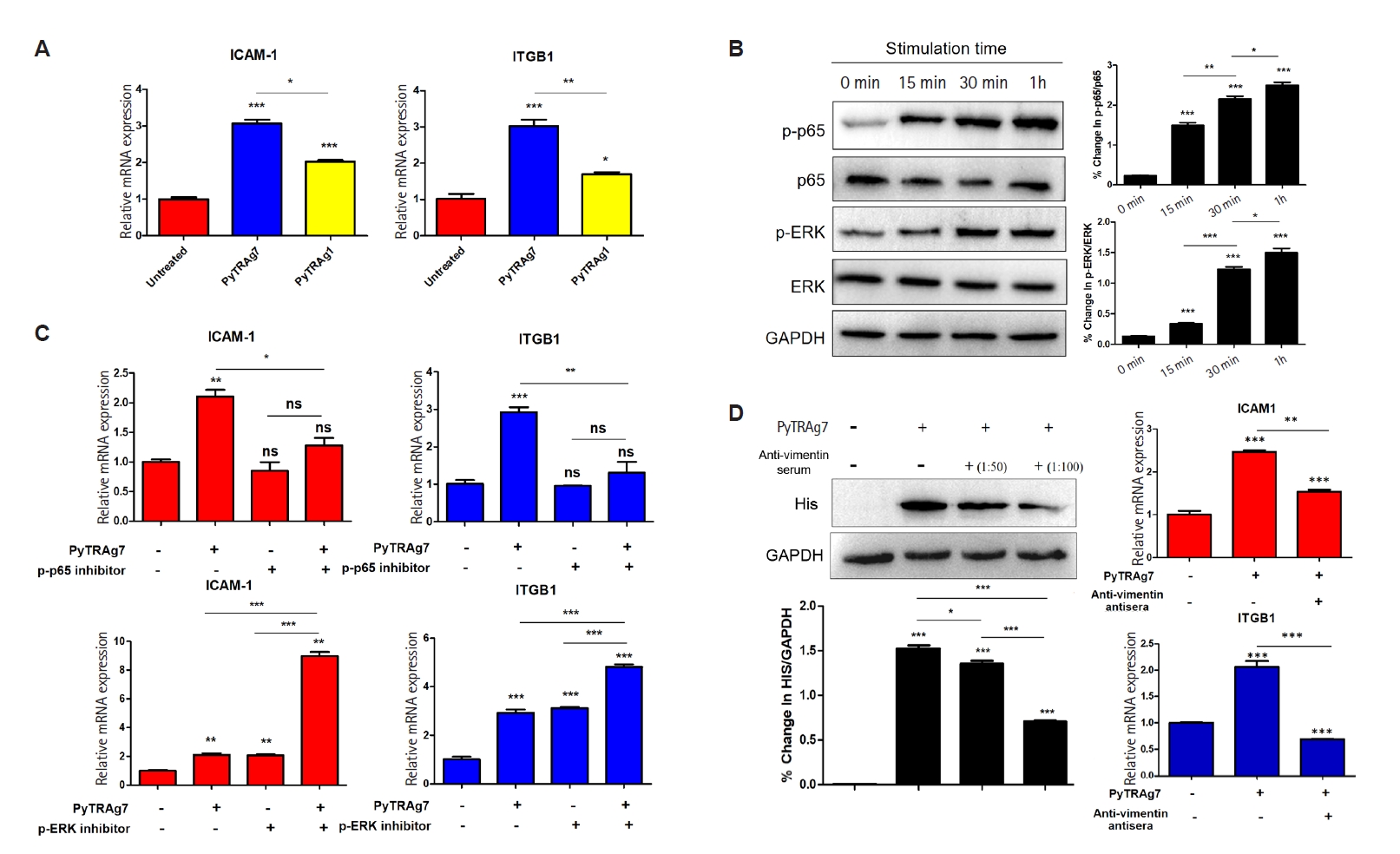
Fig. 4.Attenuated virulence and reduced splenic accumulation of ΔTRAg7 parasites in mice. (A) The parasitemia of ΔTRAg7 strain was less than the wild type. BALB/c mice were infected with 1×106 ΔTRAg7 or wild-type (WT) parasites; blood smears from day 3 post-infection onward were Giemsa-stained to quantify parasitemia. (B) The survival rate of mice infected with ΔTRAg7 strain was improved. From the first day of infection, the survival rate of mice in both groups was measured. (C) Comparison of spleen-body ratio in mice. (D,E) Changes of spleens’ adhesion molecules in mice infected with different strains. The protein levels (D) and mRNA levels (E) of ICAM-1 and ITGB1 in the spleens of mice infected with Plasmodium were detected by Western blot and qPCR. (F) Ex vivo adhesion of infected red blood cells (iRBCs) to mouse splenic fibroblasts (MSFs). Left: representative micrographs (400× and 1,000×; arrows indicate adherent iRBCs). Right: quantification of mean adherent iRBCs per MSF (mean±SD; n≥3 biological replicates; ≥150 MSFs counted per replicate). (G,H) The parasites in spleen of mice in ΔTRAg7 group was significantly lower than that in WT group. Hematoxylin-eosin staining of spleens (G) showed ΔTRAg7 group’s structures are relatively complete, and the degree of malarial pigment deposition was obviously light. The spleen of 3 groups of mice was taken to detect the DNA level of Plasmodium (H), showed that the parasites charge in spleen of mice in ΔTRAg7 group was significantly lower. Statistics: one-way ANOVA with Dunnett’s test (nsP≥0.05; *P<0.05, **P<0.01, ***P<0.001).
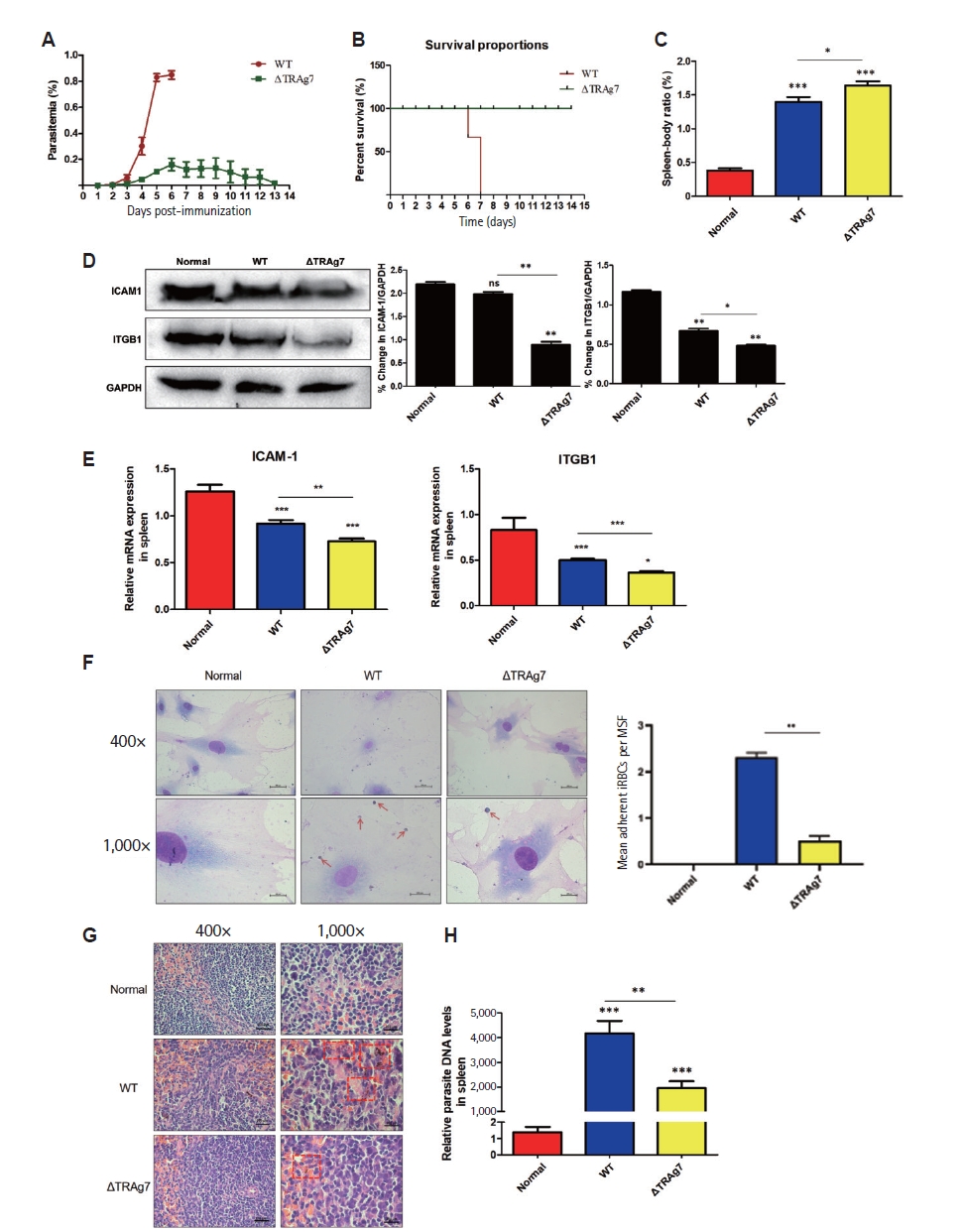
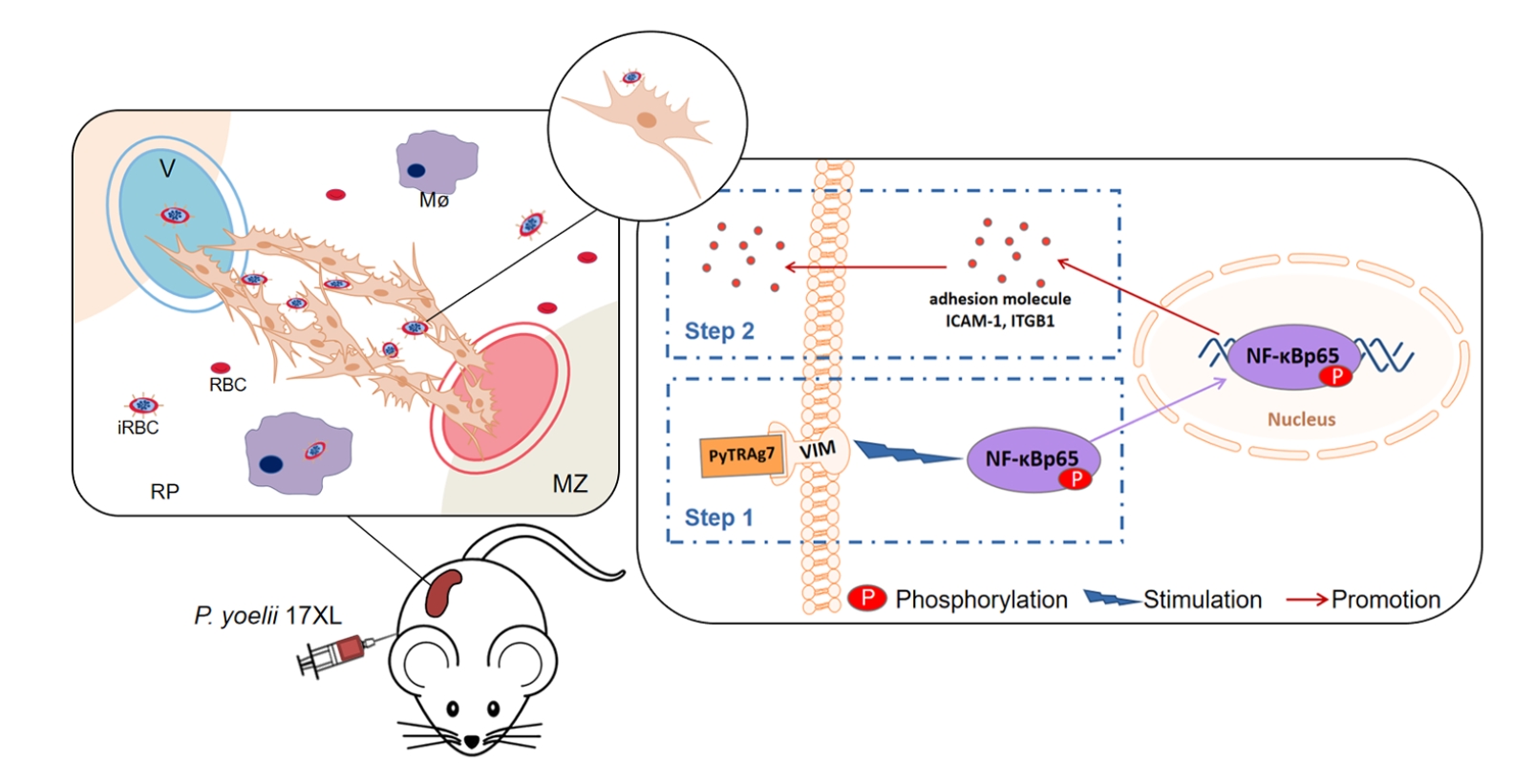
Fig. 5.Proposed model of PyTRAg7-mediated immune evasion in the spleen during Plasmodium yoelii infection. During the intraerythrocytic stage, PyTRAg7 is exported to the surface of infected red blood cells (iRBCs). It binds specifically to vimentin on the surface of mouse splenic fibroblasts (MSFs). This interaction activates the NF-κB p65 pathway, leading to upregulation of adhesion molecules ICAM-1 and ITGB1 in MSFs. The elevated expression of adhesion molecules enhances iRBCs adherence to MSFs and other splenic barrier cells. These cells encapsulate the parasites and spatially segregate them from immune effector cells, thereby facilitating immune evasion. This sequestration allows parasites to bypass splenic clearance and transit into peripheral circulation to continue the infection cycle. V, venous lumen; Mø, macrophages; RP, red pulp; MZ, marginal zone.
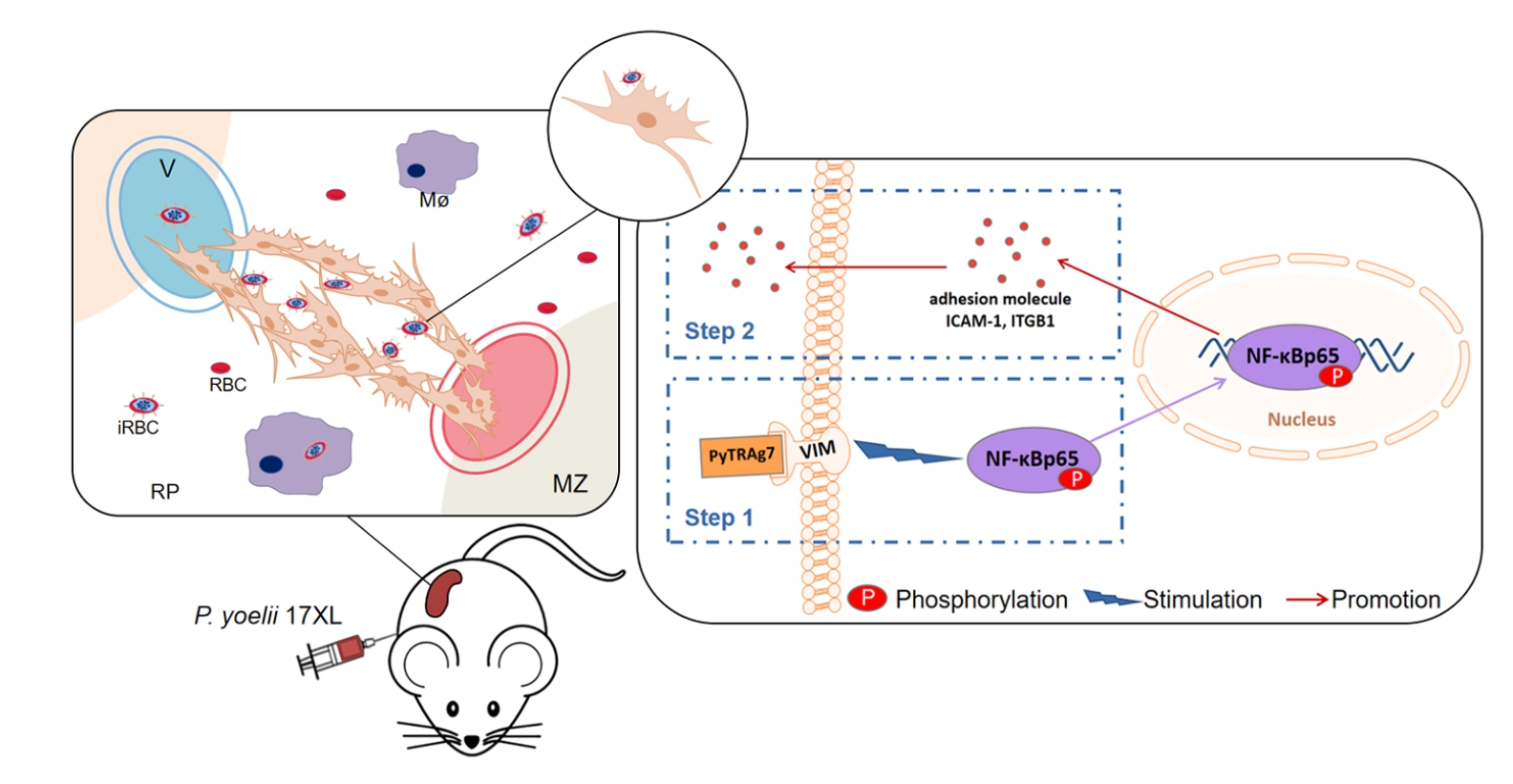
Table 1.The results of silver stain gel identified by mass spectrometry
Table 1.
|
Protein No. |
Accession No. |
Protein name |
Coverage (%) |
Molecular weight (kDa) |
Peptide identified |
|
1 |
P20152 |
Vimentin |
44.0 |
53.557 |
16 |
|
2 |
Q3V235 |
Prohibitin-2 |
16.7 |
33.296 |
8 |
|
3 |
P68040 |
Guanine nucleotide-binding protein subunit beta-2-like 1 |
12.0 |
35.076 |
3 |
|
4 |
Q8BMK4 |
Cytoskeleton-associated protein 4 |
4.3 |
63.691 |
2 |
|
5 |
Q8VDD5 |
Myosin-9 |
7.1 |
226.37 |
2 |
|
6 |
Q91ZX7 |
Low-density lipoprotein receptor-related protein 1 |
0.7 |
504.74 |
2 |
References
- 1. World Health Organization. World malaria report 2024. Geneva. World Health Organization; 2024.
- 2. Dondorp AM, Ince C, Charunwatthana P, et al. Direct in vivo assessment of microcirculatory dysfunction in severe falciparum malaria. J Infect Dis 2008;197:79-84. https://doi.org/10.1086/523762
- 3. Schofield L. Intravascular infiltrates and organ-specific inflammation in malaria pathogenesis. Immunol Cell Biol 2007;85:130-7. https://doi.org/10.1038/sj.icb.7100040
- 4. Avril M, Brazier AJ, Melcher M, Sampath S, Smith JD. DC8 and DC13 var genes associated with severe malaria bind avidly to diverse endothelial cells. PLoS Pathog 2013;9:e1003430. https://doi.org/10.1371/journal.ppat.1003430
- 5. Baruch DI, Pasloske BL, Singh HB, et al. Cloning the P. falciparum gene encoding PfEMP1, a malarial variant antigen and adherence receptor on the surface of parasitized human erythrocytes. Cell 1995;82:77-87. https://doi.org/10.1016/0092-8674(95)90054-3
- 6. Zhang H, Shen F, Yu J, et al. Plasmodium vivax protein PvTRAg23 triggers spleen fibroblasts for inflammatory profile and reduces type I collagen secretion via NF-kappaBp65 pathway. Front Immunol 2022;13:877122. https://doi.org/10.3389/fimmu.2022.877122
- 7. Kong WZ, Zhang HY, Sun YF, et al. Plasmodium vivax tryptophan-rich antigen reduces type I collagen secretion via the NF-kappaBp65 pathway in splenic fibroblasts. Parasit Vectors 2024;17:239. https://doi.org/10.1186/s13071-024-06264-y
- 8. Engwerda CR, Beattie L, Amante FH. The importance of the spleen in malaria. Trends Parasitol 2005;21:75-80. https://doi.org/10.1016/j.pt.2004.11.008
- 9. Buffet PA, Safeukui I, Milon G, Mercereau-Puijalon O, David PH. Retention of erythrocytes in the spleen: a double-edged process in human malaria. Curr Opin Hematol 2009;16:157-64. https://doi.org/10.1097/MOH.0b013e32832a1d4b
- 10. Lewis SM, Williams A, Eisenbarth SC. Structure and function of the immune system in the spleen. Sci Immunol 2019;4:eaau6085. https://doi.org/10.1126/sciimmunol.aau6085
- 11. Fernandez-Becerra C, Bernabeu M, Castellanos A, et al. Plasmodium vivax spleen-dependent genes encode antigens associated with cytoadhesion and clinical protection. Proc Natl Acad Sci U S A 2020;117:13056-65. https://doi.org/10.1073/pnas.1920596117
- 12. Gowda DC, Wu X. Parasite recognition and signaling mechanisms in innate immune responses to malaria. Front Immunol 2018;9:3006. https://doi.org/10.3389/fimmu.2018.03006
- 13. Tuikue Ndam N, Moussiliou A, Lavstsen T, et al. Parasites causing cerebral falciparum malaria bind multiple endothelial receptors and express EPCR and ICAM-1-binding PfEMP1. J Infect Dis 2017;215:1918-25. https://doi.org/10.1093/infdis/jix230
- 14. Fu H, Lu J, Zhang X, et al. Identification of the recombinant Plasmodium vivax surface-related antigen as a possible immune evasion factor against human splenic fibroblasts by targeting ITGB1. Front Cell Dev Biol 2021;9:764109. https://doi.org/10.3389/fcell.2021.764109
- 15. Ntumngia FB, Bouyou-Akotet MK, Uhlemann AC, et al. Characterisation of a tryptophan-rich Plasmodium falciparum antigen associated with merozoites. Mol Biochem Parasitol 2004;137:349-53. https://doi.org/10.1016/j.molbiopara.2004.06.008
- 16. Jalah R, Sarin R, Sud N, et al. Identification, expression, localization and serological characterization of a tryptophan-rich antigen from the human malaria parasite Plasmodium vivax. Mol Biochem Parasitol 2005;142:158-69. https://doi.org/10.1016/j.molbiopara.2005.01.020
- 17. Zeeshan M, Tyagi RK, Tyagi K, Alam MS, Sharma YD. Host-parasite interaction: selective Pv-fam-a family proteins of Plasmodium vivax bind to a restricted number of human erythrocyte receptors. J Infect Dis 2015;211:1111-20. https://doi.org/10.1093/infdis/jiu558
- 18. Burns JM, Adeeku EK, Belk CC, Dunn PD. An unusual tryptophan-rich domain characterizes two secreted antigens of Plasmodium yoelii-infected erythrocytes. Mol Biochem Parasitol 2000;110:11-21. https://doi.org/10.1016/s0166-6851(00)00252-8
- 19. De Niz M, Heussler VT. Rodent malaria models: insights into human disease and parasite biology. Curr Opin Microbiol 2018;46:93-101. https://doi.org/10.1016/j.mib.2018.09.003
- 20. Pattaradilokrat S, Cheesman SJ, Carter R. Congenicity and genetic polymorphism in cloned lines derived from a single isolate of a rodent malaria parasite. Mol Biochem Parasitol 2008;157:244-7. https://doi.org/10.1016/j.molbiopara.2007.10.011
- 21. Jaijyan DK, Singh H, Singh AP. A sporozoite- and liver stage-expressed tryptophan-rich protein plays an auxiliary role in Plasmodium liver stage development and is a potential vaccine candidate. J Biol Chem 2015;290:19496-511. https://doi.org/10.1074/jbc.M114.588129
- 22. Wu Y, He J, Cao X, et al. Progress of researches on mechanisms underlying immune escape of Plasmodium. Zhongguo Xue Xi Chong Bing Fang Zhi Za Zhi 2025;37:325-31. https://doi.org/10.16250/j.32.1915.2024214
- 23. Su XZ, Xu F, Stadler RV, Teklemichael AA, Wu J. Malaria: factors affecting disease severity, immune evasion mechanisms, and reversal of immune inhibition to enhance vaccine efficacy. PLoS Pathog 2025;21:e1012853. https://doi.org/10.1371/journal.ppat.1012853
- 24. Aparici Herraiz I, Caires HR, Castillo-Fernández Ó, et al. Advancing key gaps in the knowledge of Plasmodium vivax cryptic infections using humanized mouse models and organs-on-chips. Front Cell Infect Microbiol 2022;12:920204. https://doi.org/10.3389/fcimb.2022.920204
- 25. Mirzadegan T, Benkö G, Filipek S, Palczewski K. Sequence analyses of G-protein-coupled receptors: similarities to rhodopsin. Biochemistry 2003;42:2759-67. https://doi.org/10.1021/bi027224+
- 26. Loewenstein Y, Raimondo D, Redfern O C, et al. Protein function annotation by homology-based inference. Genome Biol 2009;10:207. https://doi.org/10.1186/gb-2009-10-2-207
- 27. Qian WJ, Yan JS, Gang XY, et al. Intercellular adhesion molecule-1 (ICAM-1): from molecular functions to clinical applications in cancer investigation. Biochim Biophys Acta Rev Cancer 2024;1879:189187. https://doi.org/10.1016/j.bbcan.2024.189187
- 28. Virtanen I, Korhonen M, Kariniemi A L, et al. Integrins in human cells and tumors. Cell Differ Dev 1990;32:215-27. https://doi.org/10.1016/0922-3371(90)90034-t
- 29. Alieva IB, Shakhov AS, Dayal AA, et al. Unique role of vimentin in the intermediate filament proteins family. Biochemistry (Mosc) 2024;89:726-36. https://doi.org/10.1134/S0006297924040114
- 30. Suprewicz Ł, Swoger M, Gupta S, et al. Extracellular vimentin as a target against SARS-CoV-2 host cell invasion. Small 2022;18:e2105640. https://doi.org/10.1002/smll.202105640
- 31. Feng X, Yu JL, Sun YF, et al. Plasmodium yoelii surface-related antigen (PySRA) modulates the host pro-inflammatory responses via binding to CD68 on macrophage membrane. Infect Immun 2024;92:e0011324. https://doi.org/10.1128/iai.00113-24
- 32. Leoni S, Buonfrate D, Angheben A, Gobbi F, Bisoffi Z. The hyper-reactive malarial splenomegaly: a systematic review of the literature. Malar J 2015;14:185. https://doi.org/10.1186/s12936-015-0694-3
- 33. Ghosh D, Stumhofer JS. The spleen: "epicenter" in malaria infection and immunity. J Leukoc Biol 2021;110:753-69. https://doi.org/10.1002/JLB.4RI1020-713R
- 34. Lacerda-Queiroz N, Riteau N, Eastman RT, et al. Mechanism of splenic cell death and host mortality in a Plasmodium yoelii malaria model. Sci Rep 2017;7:10438. https://doi.org/10.1038/s41598-017-10776-2
- 35. Boullé C, Lebredonchel E, Campillo JT, et al. Association between Loa loa microfilaremia and anatomical hyposplenia in a rural area of the Republic of Congo: a population-based cross-sectional study. Infect Dis Poverty 2025;14:8. https://doi.org/10.1186/s40249-025-01277-w