Abstract
Acanthamoeba is a genus of free-living amoebae commonly found in soil, water, and other habitats. This organism undergoes 2 distinct stages in its life cycle, the trophozoite and the cyst. Under adverse conditions, trophozoites transform into cysts, which are notably resistant to harsh physical and chemical conditions. Infection by Legionella pneumophila has been shown to decrease the number of cysts in its host Acanthamoeba species, although the mechanisms responsible for this effect remain poorly understood. In this study, A. castellanii was co-cultured with either L. pneumophila or Escherichia coli to assess the impact on encystation and to explore the genes involved in this process. Following a 72-h encystation induction period, it was observed that Acanthamoeba infected with Legionella exhibited a 45.8% reduction in cyst formation compared to the control group. In contrast, Acanthamoeba that phagocytosed E. coli showed a 21.7% decrease. To identify the genes involved in this phenomenon, real-time PCR analysis was conducted on 20 genes known to be upregulated during encystation. This analysis was performed to verify their expression patterns at 24, 48, and 72 h. Notably, ten genes, including cyst-specific protein 21, glycosyltransferase, RSNARE, and cellulose synthase, did not exhibit increased expression in Legionella-infected Acanthamoeba. However, these genes showed elevated expression levels in both the control group and the bacteria-phagocytosed Acanthamoeba. This suggests that several cellular processes, including cell wall formation, are inhibited in Acanthamoeba infected with Legionella, resulting in reduced encystation.
-
Key words: Acanthamoeba castellanii, Legionella pneumophila, encystation
Introduction
Acanthamoeba spp. are ubiquitous free-living amoebae prevalent in diverse aquatic environments, capable of causing significant human diseases such as amoebic keratitis and granulomatous amoebic encephalitis [
1]. These amoebae exist in 2 distinct forms: the motile, feeding trophozoite stage and the dormant, resistant cyst stage [
2]. While the trophozoite represents the active form, the cyst serves as a vital survival mechanism, especially under harsh environmental conditions. This encystation process is crucial for their persistence in extreme habitats. Beyond their role as passive environmental inhabitants,
Acanthamoeba spp. serve as hosts for various pathogenic bacteria.
Acanthamoeba are often referred to as the ‘Trojan horse’ of microorganisms including
Escherichia coli O157,
Legionella pneumophila,
Coxiella burnetii,
Helicobacter pylori,
Chlamydophila pneumoniae,
Vibrio cholerae,
Listeria monocytogenes,
Campylobacter jejuni,
Mycobacterium leprae and
Pseudomonas aeruginosa [
3-
5].
Among these,
L. pneumophila, the causative agent of Legionnaires’ disease, is an intracellular pathogen that exploits protozoa including
Acanthamoeba, as reservoirs and vectors for transmission to humans [
6].
A. castellanii can harbor
L. pneumophila within their cells, provides a protective niche, shielding them from disinfection processes. Despite this symbiotic protection, the presence of
L. pneumophila exerts a profound impact on the biology of
Acanthamoeba, notably inhibiting DNA replication and cell division, which ultimately leads to the arrest in the proliferation of the host [
7,
8].
Legionella infection disrupts the normal life cycle of
Acanthamoeba, particularly affecting the encystation process by altering cellular proliferation, morphology, and gene expression linked to encystment. Notably,
Legionella amylase A (LamA)-mediated glycogenolysis depletes cytosolic glycogen reserves and hinders the synthesis of the cellulose-rich cyst wall, effectively subverting the encystation process [
9]. Despite emerging evidence of
Legionella’s impact on
Acanthamoeba encystation, the precise molecular mechanisms remain inadequately understood. Specifically, understanding how
L. pneumophila infection modulates the gene expression associated with encystation could provide valuable insights into the survival strategies of both the
Acanthamoeba and the
Legionella.
This study aims to elucidate the effects of
L. pneumophila infection on the encystation of
A. castellanii. Previous research has identified essential genes involved in the encystation of
A. castellanii through mRNA sequencing and real-time PCR analysis [
10-
15]. Building on this knowledge, we compare the encystation rates and gene expression profiles of
A. castellanii infected with
L. pneumophila to those of amoebae that have phagocytosed non-pathogenic
E. coli. Through this comparison, we seek to unravel the molecular mechanisms underlying these interactions. Our findings will enhance the understanding of amoebal-bacterial interactions and may inform future strategies for controlling
Legionella outbreaks and
Acanthamoeba encystation in the environmental contexts.
Methods
Ethics statement
Not applicable.
Cell cultures
A. castellanii (ATCC 30868) was cultured axenically in peptone-yeast-glucose (PYG) media (20 g proteose peptone, 1 g yeast extract, 0.1 M glucose, 4 mM MgSO4, 0.4 mM CaCl2, 3.4 mM sodium citrate, 0.05 mM Fe(NH4)2(SO4)2, 2.5 mM Na2HPO4, and 2.5 mM K2HPO4 in 1 L distilled water with the final pH adjusted to 6.5) at 25°C, and L. pneumophila (ATCC 33152) was cultured on a buffered charcoal yeast extract agar plate (Thermo Fisher Scientific) at 37°C with 5% CO2. E. coli (Enzynomics) was cultured in tryptone-yeast-NaCl (Luria–Bertani) media at 37°C using a shaking incubator.
Infection of bacteria
L. pneumophila was diluted in PBS until the OD
600 absorbance reading reached 1, which corresponds to 10
9 CFU/ml. A total of 1×10
7 A. castellanii trophozoites were incubated with 1 ml of the
Legionella suspension at 37℃ with 5% CO
2 for 1 h in PYG medium. After incubation,
Acanthamoeba was washed with Page’s amoeba saline and then incubated with fresh PYG media containing 100 μg/ml of gentamicin for 2 h to kill any extracellular
Legionella. The
Acanthamoeba infected with
Legionella (A+L) were washed with Page’s amoeba saline twice and incubated with fresh PYG media for 12 h in a 25℃ incubator.
Acanthamoeba (control) or those that had phagocytosed
E. coli (A+E) were treated in the same way [
16].
Encystation of Acanthamoeba
Encystation of
A. castellanii (control, A+E, and A+L) was induced using an encystment media composed of 95 mM NaCl, 5 mM KCl, 8 mM MgSO
4, 0.4 mM CaCl
2, 1 mM NaHCO
3, 20 mM Tris-HCl at pH 9.0. The encystation process for each group was monitored at 24, 48, and 72 h, with morphological changes observed using light microscopy. Mature cysts were counted under a microscope after treatment with 0.5% SDS to calculate encystation ratios [
10,
17].
The expression of target genes was determined by real-time PCR analysis. The total RNA was purified using an RNeasy Mini kit (Qiagen), and the cDNA was synthesized using a RevertAid first-strand cDNA synthesis kit (Thermo Fisher Scientific) following the manufacturer’s instructions. Real-time PCR was conducted using a Magnetic Induction Cycler PCR machine (PhileKorea) as manufacturer’s instructions: preincubation at 95°C for 1 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 30 sec. All reaction mixtures were made using a Luna Universal qPCR Master Mix (New England Biolabs) with different sense and antisense primers (
Table 1). The relative expression levels were calculated by normalizing the critical threshold (Ct) values to that of the internal control (18S rDNA), and graphs were presented using the 2^-ΔΔCt method [
18].
All statistical analyses were conducted using GraphPad Prism version 8.0. Data are presented as the mean±SD. Statistical significance between groups is denoted by asterisks, with P-values less than 0.05 considered statistically significant (*P<0.05, **P<0.01).
Results
Inhibition of encystation of A. castellanii by L. pneumophila infection
To investigate the effect of
Legionella infection on the encystation of
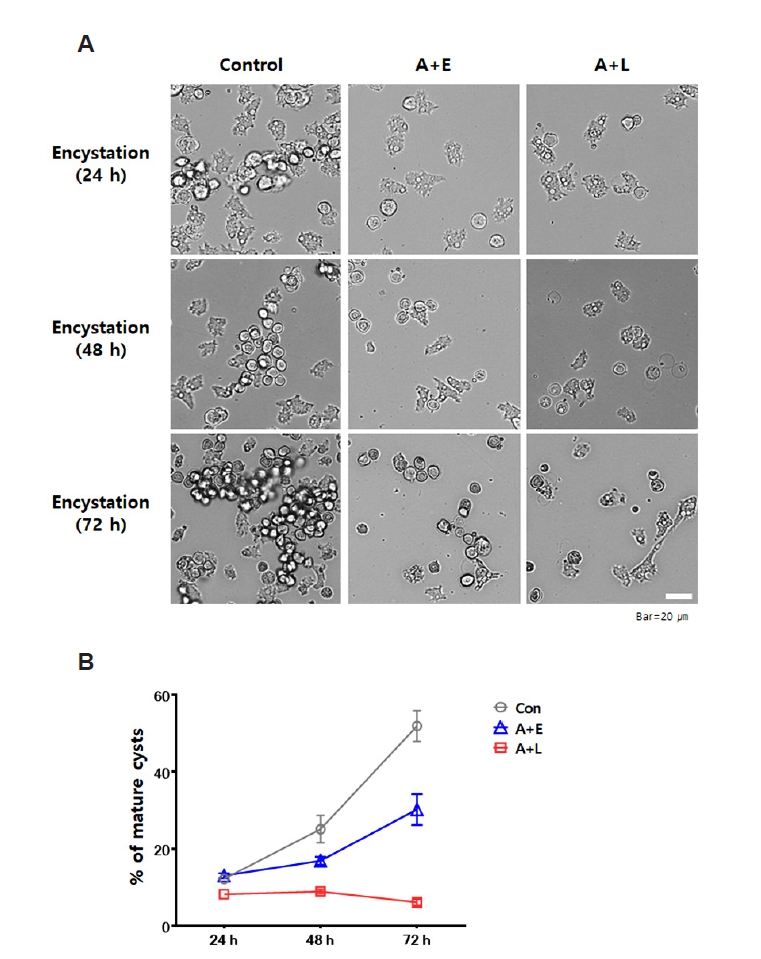
Acanthamoeba, cyst formation experiments were conducted using several groups: control, A+E, and A+L. Encystation was induced over a period of 72 h. Encystation of each group was monitored at 24, 48, and 72 h using microscopy (
Fig. 1A), and the number of mature cysts (%) was graphically represented in
Fig. 1B.
During the encystation induction periods of 24, 48, and 72 h, the control group showed encystment rates of 12.2%, 25.1%, and 51.9%, respectively. In contrast, the A+L group demonstrated encystation rates of 8.2%, 8.9%, and 6.1% over the same time intervals, and the A+E group showed rates of 13.0%, 16.9%, and 30.2%. The A+L group exhibited a significant reduction in encystment, with a 45.8% decrease compared to the control group (from 51.9% to 6.1%, corresponding to an 88% relative decrease), and the A+E group showed a 21.7% reduction in encystment relative to the control (from 51.9% to 30.2%, corresponding to a 42% relative decreases). These findings suggest that the presence of L. pneumophila or E. coli negatively impacts the encystation process of A. castellanii, with L. pneumophila infection significantly reducing encystment more than E. coli.
Gene expression analysis during encystation
To further investigate the molecular mechanisms underlying the encystation of
Acanthamoeba infected by
Legionella, real-time PCR analysis was conducted on 20 genes previously identified as upregulated during
Acanthamoeba encystation [
10-
15]. In previous mRNA sequencing analysis of
A. castellanii results, 13 genes that showed a significant increase in expression 24 h after encystation induction are summarized in
Table 2 [
10]. Additionally, 7 genes that showed increased expression related to autophagy or cyst wall formation during the encystation process of
A. castellanii are also summarized in
Table 2 [
11-
15]. This study observed changes in the expression of these 20 genes when
A. castellanii formed cysts after being infected with
L. pneumophila or after phagocytosing
E. coli (
Supplementary Fig. S1).
In
A. castellanii infected with
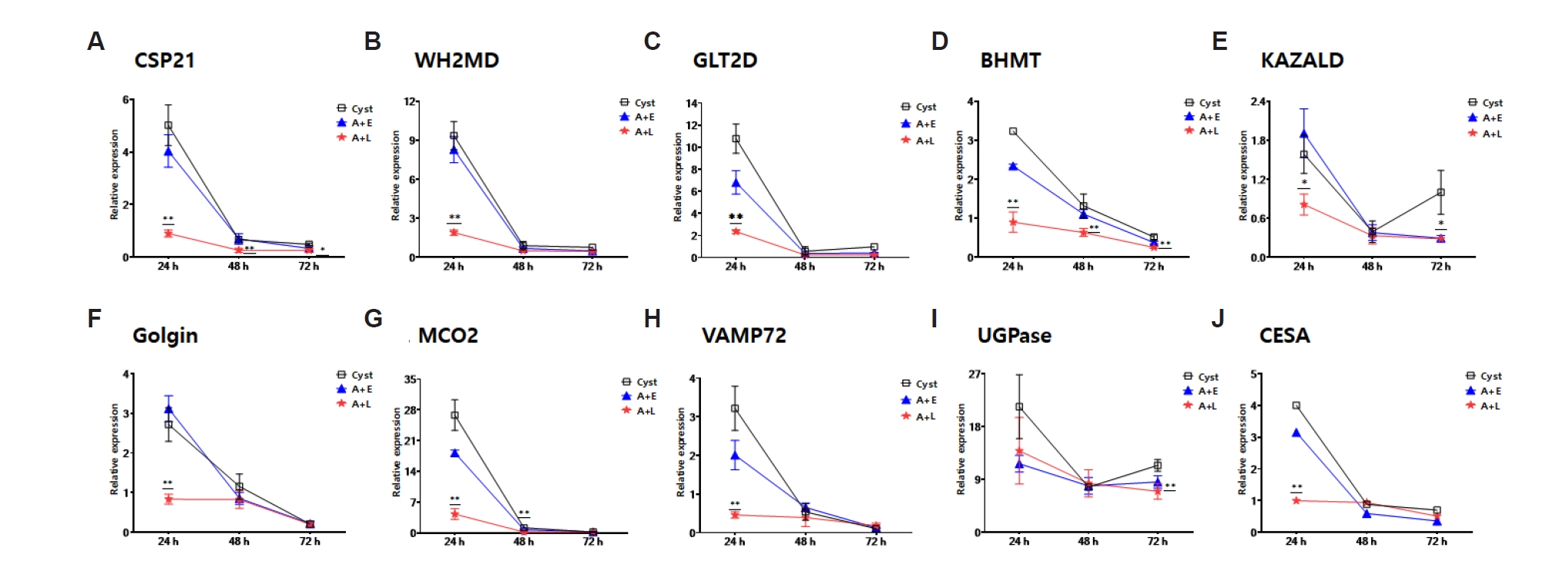
L. pneumophila, 10 out of the 20 genes were downregulated in 24 h encysting cells compared to the control group, and remained decreased until 72 h (
Fig. 2). These genes include key proteins such as the cyst-specific protein 21 (
Fig. 2A), WH2 motif domain-containing protein (
Fig. 2B), glycosyltransferase (
Fig. 2C), betaine homocysteine methyltransferase (
Fig. 2D), kazal type serine protease inhibitor domain containing protein (
Fig. 2E), golgi family protein (
Fig. 2F), multicopper oxidase (
Fig. 2G), RSNARE (
Fig. 2H), UDP-glucose pyrophosphorylase (
Fig. 2I), and cellulose synthase (
Fig. 2J) involved in cellular and metabolic processes. The functions of these ten genes seem to be primarily involved in cyst wall formation (
Table 3). The lack of upregulation of these genes suggests that
L. pneumophila may inhibit or alter the normal encystation-related structure components, potentially contributing to the observed reduction in encystment.
Whereas cysteine proteinases and encystation-mediating serine proteinase were increased at 48 and 72 h, and other encystation-related genes (such as twin arginine translocation pathway protein, aminotransferase, von Willebrand factor type A domain containing protein, armadillo/beta catenin-like repeat domain containing protein, autophagy proteins, encystation-mediating cysteine proteinase, and glycogen phosphorylase) were upregulated in
Legionella-infected
Acanthamoeba compared to the control group (
Supplementary Fig. S1).
Discussion
This study provides significant insights into the interaction between A. castellanii and L. pneumophila, specifically focusing on how Legionella infection affects the encystation process of Acanthamoeba. Our findings reveal that L. pneumophila not only significantly reduces encystation rates of Acanthamoeba but also disrupts the gene expression program necessary for encystation.
The encystation process is a critical adaptive mechanism for
Acanthamoeba, allowing it to persist in harsh environmental conditions. Our results showed a marked reduction in encystation rates in
A. castellanii infected with
L. pneumophila compared to the control group (
Fig. 1). This suggests that
L. pneumophila infection compromises the ability of
Acanthamoeba to form protective cysts, potentially increasing the amoeba’s susceptibility to environmental stressors. If the encystment of
Acanthamoeba is inhibited, it also means that its ability to protect the internal
Legionella from environmental threats is reduced. It is worth considering why
Legionella would inhibit the encystment of its protective host,
Acanthamoeba. The reduction in encystation was more pronounced with
L. pneumophila (A+L) than with non-pathogenic
E. coli (A+E), indicating a pathogen-specific interaction. The encystation media induce cyst formation in
Acanthamoeba due to nutrient deprivation. In the A+E group, however, the phagocytosis and digestion of
E. coli provide nutrients, which is believed to result in a reduced encystation rate.
Gene expression analysis further supported these findings, demonstrating that
L. pneumophila alters the expression of key genes involved in the encystation pathway. Notably, 10 out of 20 genes typically upregulated during encystation did not show increased expression in the presence of
L. pneumophila (
Fig. 2). This includes genes encoding cyst-specific proteins (
Fig. 2A) and enzymes essential for cyst wall biosynthesis, such as UDP-glucose pyrophosphorylase and cellulose synthase (
Fig. 2I,
J). Previous studies have shown that
L. pneumophila can manipulate host cells by degrading cellular glycogen reserves [
9]. This process involves
Legionella amylase A in glycogenolysis, thereby hindering cyst wall formation. In
Giardia lamblia, RSNAREs are involved in the encystation pathway, where they contribute to the formation of the cyst wall, a protective structure composed of cyst wall material [
19]. Similarly, in
Entamoeba, RSNAREs play a role in the trafficking of cyst wall components [
20]. However, in
L. pneumophila-infected
A. castellanii, the expression of Golgi family proteins and RSNARE is notably diminished (
Fig. 2F,
H). The absence of upregulation in these genes suggests that
L. pneumophila may interfere with the regulatory pathways that initiate and sustain the cyst wall formation during the encystation process. The mechanism by which
L. pneumophila exerts these effects may involve the depletion of resources or direct modulation of host cellular pathways.
The transition of
Acanthamoeba into the cyst form typically involves the upregulation of cysteine and serine proteinase for autophagy and internal digestion [
13,
14]. These processes were found to be suppressed at 24 h by
L. pneumophila infection (
Supplementary Fig. S1P,
Q). However, these 2 genes were observed to increase again at 48 or 72 h. This is presumed to be because
Legionella delays the delivery to lysosomes in host cells, thereby inhibiting autophagy formation for its survival [
21]. Meanwhile, other encystation-related genes (such as twin arginine translocation pathway protein, aminotransferase, von Willebrand factor type A domain containing protein, armadillo/beta catenin-like repeat domain containing protein, autophagy proteins, encystation-mediating cysteine proteinase, and glycogen phosphorylase) were found to increase in
Acanthamoeba infected with
Legionella compared to the control group (
Supplementary Fig. S1). Further research is needed to understand the reasons and mechanisms behind this.
These findings have broader implications for understanding amoebal-bacterial interactions and the survival strategies employed by intracellular pathogens. The ability of L. pneumophila to inhibit encystation could facilitate its persistence and transmission in the environment, as Acanthamoeba serves as a reservoir and vector for the bacteria. This interaction highlights the complex ecological relationships between free-living amoebae and pathogenic bacteria, emphasizing the need for further research to uncover the precise molecular mechanisms involved. Future studies should focus on elucidating the signaling pathways disrupted by L. pneumophila and identifying potential targets for intervention. Understanding these interactions at a molecular level could inform strategies to prevent Legionella outbreaks by targeting its amoebal hosts and disrupting its life cycle within these protozoa. Additionally, since these findings are based on a single amoeba strain (A. castellanii) and bacterial isolate (L. pneumophila), they cannot be generalized. Therefore, further research involving various amoeba strains and bacterial isolates is considered necessary.
In conclusion, L. pneumophila appears to significantly inhibit the cyst formation of A. castellanii by suppressing the expression of genes involved in cyst formation, particularly those related to cyst wall formation. This study advances our understanding of how Legionella influences the encystation of Acanthamoeba.
Notes
-
Author contributions
Conceptualization: Moon EK. Data curation: Lee HA, Quan FS, Kong HH, Moon EK. Funding acquisition: Moon EK. Investigation: Jo HJ, Lee HA, Moon EK. Methodology: Jo HJ, Lee HA, Quan FS, Kong HH, Moon EK. Validation: Moon EK. Writing – original draft: Jo HJ. Writing – review & editing: Lee HA, Quan FS, Kong HH, Moon EK.
-
Conflict of interest
Fu-Shi Quan serves as an editor of Parasites, Hosts and Diseases but had no involvement in the decision to publish this article. No other potential conflicts of interest relevant to this study were reported.
-
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (RS-2024-00346635).
Supplementary information
Fig. 1.Encystation rate of Acanthamoeba castellanii over 72 h. The encystation rates of A. castellanii (control) and those co-cultured with either Legionella pneumophila or Escherichia coli (A+L or A+E) were assessed by inducing encystation in encystation media over a period of 72 h. The extent of encystation was observed under a microscope at intervals of 24, 48, and 72 h (A), and the data were represented in a graph (B). The A+L group exhibited a 45.8% reduction in encystment, while the A+E group showed a 21.7% decrease compared to the control group.
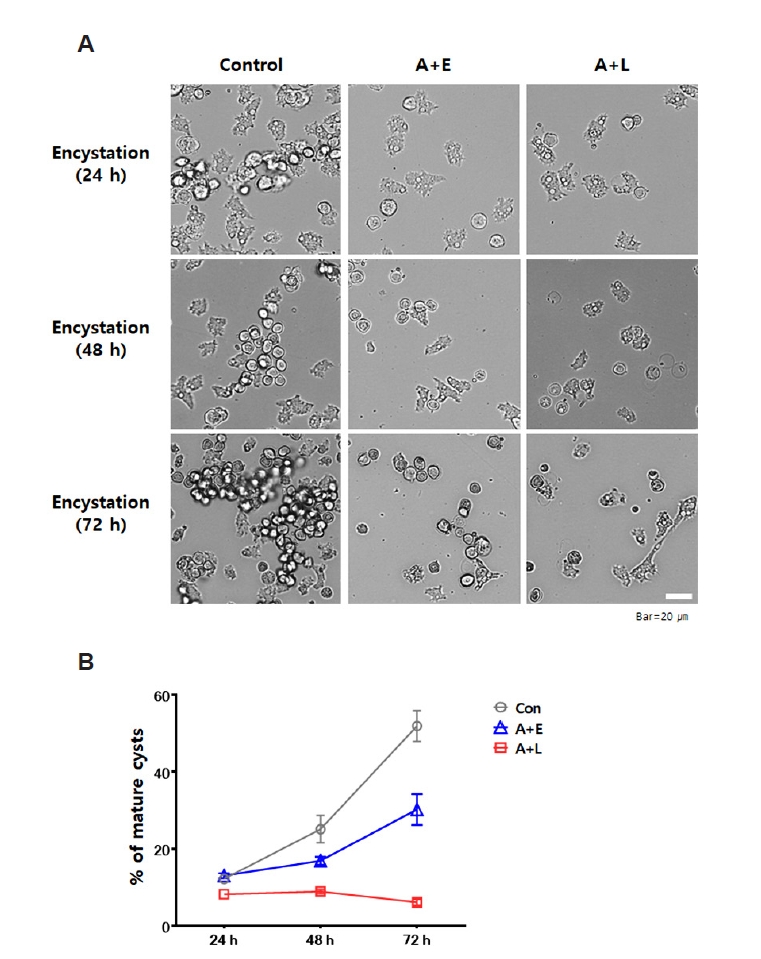
Fig. 2.Real-time PCR analysis of genes related to encystation. The expression changes of 10 out of 20 genes, previously identified to be upregulated during encystation of Acanthamoeba, were observed to be downregulated when A. castellanii formed cysts after being infected with Legionella pneumophila or after phagocytosing Escherichia coli. (A) Cyst specific protein 21 (CSP21), (B) WH2 motif domain containing protein (WH2MD), (C) glycosyltransferase, group 2 domain containing protein (GLT2D), (D) betaine homocysteine methyltransferase (BHMT), (E) Kazal type serine protease inhibitor domain containing protein (KAZALD), (F) Golgi family protein, putative (Golgin), (G) multicopper oxidase, type 2 (MCO2), (H) RSNARE, VAMP72-family (VAMP72), (I) UDP-glucose pyrophosphorylase (UGPase), and (J) cellulose synthase (CESA). A+E, A. castellanii that phagocytosed E. coli; A+L, A. castellanii infected with L. pneumophila.
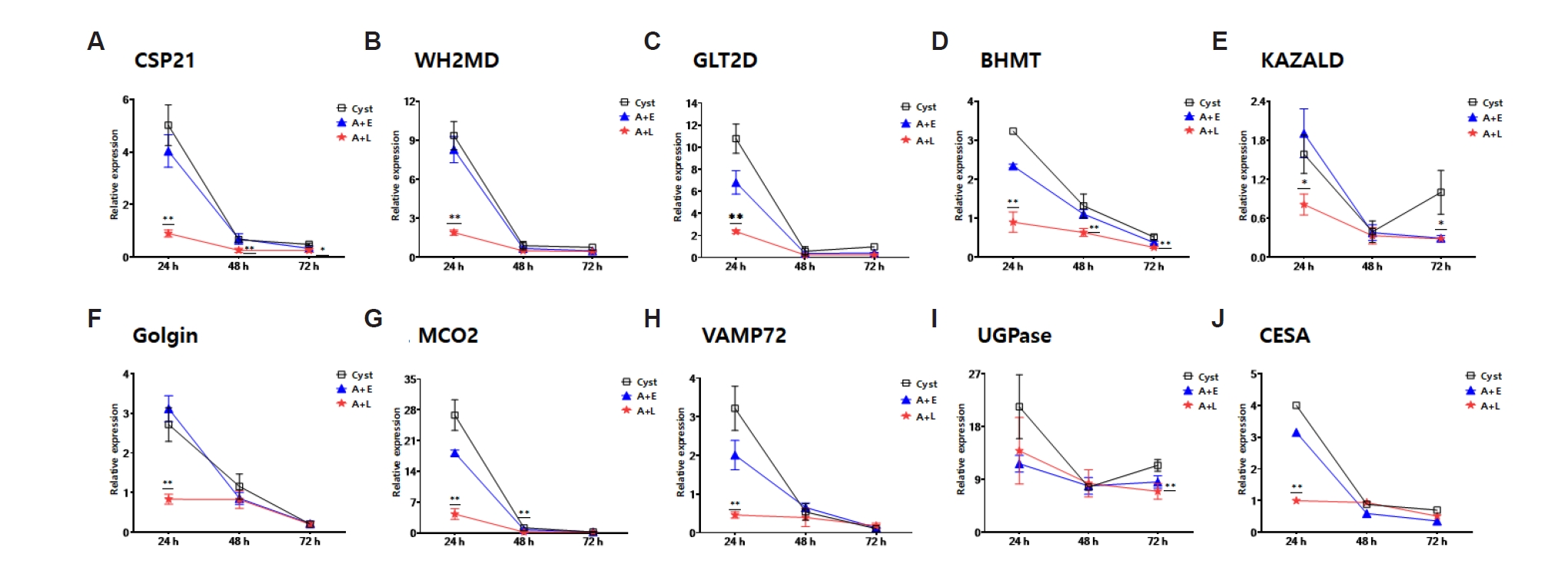
Table 1.Primer sequences for real-time PCR
Table 1.
|
Product |
Primer sequences (5’→3’) |
|
Cyst specific protein 21 |
F: gcttcctgcgcatgtgtaag |
|
R: cctccctcttgccgatgatc |
|
WH2 motif domain containing protein |
F: gaagaagagggcgagctcag |
|
R: ttgaggttgaactgctgcca |
|
Twin arginine translocation pathway protein |
F: gagaccatcacccttcgcaa |
|
R: ggatgcagtagggcctcttc |
|
Aminotransferase, class III superfamily protein |
F: ttcatgaagggtgttcgcga |
|
R: ggatgtcgggcttgatgtca |
|
Glycosyltransferase, group 2 domain containing protein |
F: acctcgagttcaagtggagc |
|
R: ttttccacgccccagatctc |
|
Betaine homocysteine methyltransferase |
F: aatctgtgggaggggaagga |
|
R: ttcggcgatgacaaagtcca |
|
Kazal type serine protease inhibitor domain containing protein |
F: cttctccgagcagctctacg |
|
R: gacgagaaggggctgtagtc |
|
Golgi family protein, putative |
F: agtcgatcgtgaagctccac |
|
R: ggcgagtcggtgaagaagaa |
|
Cysteine proteinase |
F: caccaccatcaccaccatca |
|
R: gcgtgcttcttgttccagtg |
|
von Willebrand factor type A domain containing protein |
F: caccaaccgcatcttcctct |
|
R: tgaagaagtaggcaccgctg |
|
Multicopper oxidase, type 2 |
F: tccagttcagagtgtcacgc |
|
R: gttgatctccaccagcgtca |
|
RSNARE, VAMP72-family |
F: agtggcctttgacttcctcg |
|
R: ccttcttcacgtcctccacc |
|
Armadillo/beta catenin-like repeat domain containing protein |
F: cgtccaccaccagaagaaca |
|
R: tgcaaggccaggttccatag |
|
Autophagy protein 8 |
F: aagaagttcctggtgcctgc |
|
R: ctcgtccttgtacttggcgt |
|
Autophagy protein 16 |
F: tctatgtgctgagctacgcg |
|
R: cttggaggcgcacatgattg |
|
Encystation-mediating cysteine proteinase |
F: ttcgtggaggagttcaaccg |
|
R: atctgctcctcctcgtcctt |
|
Encystation-mediating serine proteinase |
F: caactacacccaggacaccc |
|
R: gtgccggagatggtgttgta |
|
Glycogen phosphorylase |
F: gtccttctcgacgagcagaa |
|
R: cagaagctcctcctgttggg |
|
UDP-glucose pyrophosphorylase |
F: ccccgactctgtcttttccc |
|
R: tcttgtgagccgctaactgg |
|
Cellulose synthase |
F: tctacatgttctgcgccctg |
|
R: cagttgttgagcatgcggag |
|
18S rDNA |
F: cgtgctggggatagatcatt |
|
R: aaaggggagacctcacaacc |
Table 2.Genes upregulated in Acanthamoeba castellanii during encystation
Table 2.
|
Product |
GeneBank No. |
Reference |
|
Cyst specific protein 21 |
XM_004353421.1 |
[10] |
|
WH2 motif domain containing protein |
XM_004333237.1 |
[10] |
|
Twin arginine translocation pathway protein |
XM_004352737.1 |
[10] |
|
Aminotransferase, class III superfamily protein |
XM_004336305.1 |
[10] |
|
Glycosyltransferase, group 2 domain containing protein |
XM_004368209.1 |
[10] |
|
Betaine homocysteine methyltransferase |
XM_004340572.1 |
[10] |
|
Kazal type serine protease inhibitor domain containing protein |
XM_004368049.1 |
[10] |
|
Golgi family protein, putative |
XM_004341407.1 |
[10] |
|
Cysteine proteinase |
XM_004342180.1 |
[10] |
|
von Willebrand factor type A domain containing protein |
XM_004338072.1 |
[10] |
|
Multicopper oxidase, type 2 |
XM_004335693.1 |
[10] |
|
RSNARE, VAMP72-family |
XM_004334605.1 |
[10] |
|
Armadillo/beta catenin-like repeat domain containing protein |
XM_004346431.1 |
[10] |
|
Autophagy protein 8 |
EU935007.1 |
[11] |
|
Autophagy protein 16 |
FJ906697.1 |
[12] |
|
Encystation-mediating cysteine proteinase |
JQ253375.1 |
[13] |
|
Encystation-mediating serine proteinase |
EU365402.1 |
[14] |
|
Glycogen phosphorylase |
JX312797.1 |
[15] |
|
UDP-glucose pyrophosphorylase |
JX312798.1 |
[15] |
|
Cellulose synthase |
JX312799.1 |
[15] |
Table 3.Functions of encystation-related genes suppressed by Legionella pneumophila in Acanthamoeba castellanii during encystation
Table 3.
|
Product |
Function |
|
Cyst specific protein 21 |
Involved in the form of the cyst wall |
|
WH2 motif domain containing protein |
A group of proteins that bind to actin monomers, playing a crucial role in regulating the dynamic assembly and disassembly of actin filaments |
|
Glycosyltransferase |
Synthesize the glycoproteins and glycolipids required for the cyst wall |
|
Betaine homocysteine methyltransferase |
A zinc-dependent enzyme that converts betaine and homocysteine into dimethylglycine and methionine, regulating homocysteine levels |
|
Kazal type serine protease inhibitor domain containing protein |
A protein with a specific functional domain that inhibits serine proteases, wide range of biological processes |
|
Golgi family protein |
Essential for the transport and processing of cyst wall proteins |
|
Multicopper oxidase |
A specific copper-binding site within a class of enzymes that oxidize substrates using oxygen as the final electron acceptor |
|
RSNARE, VAMP72-family |
Involved in vesicle trafficking necessary for autophagy and encystation |
|
UDP-glucose pyrophosphorylase |
Produce UDP-glucose, a precursor for synthesizing cyst wall components such as cellulose or chitin |
|
Cellulose synthase |
Build the cyst wall by catalyzing the formation of cellulose microfibrils from glucose precursors |
References
- 1. Siddiqui R, Khan NA. Biology and pathogenesis of Acanthamoeba. Parasit Vectors 2012;5:6. https://doi.org/10.1186/1756-3305-5-6
- 2. Wang Y, Jiang L, Zhao Y, et al. Biological characteristics and pathogenicity of Acanthamoeba. Front Microbiol 2023;14:1147077. https://doi.org/10.3389/fmicb.2023.1147077
- 3. Winiecka-Krusnell J, Linder E. Bacterial infections of free-living amoebae. Res Microbiol 2001;152:613-9. https://doi.org/10.1016/S0923-2508(01)01240-2
- 4. Greub G, Raoult D. Microorganisms resistant to free-living amoebae. Clin Microbiol Rev 2004;17:413-33. https://doi.org/10.1128/CMR.17.2.413-433.2004
- 5. Axelsson-Olsson D, Waldenström J, Broman T, Olsen B, Holmberg M. Protozoan Acanthamoeba polyphaga as a potential reservoir for Campylobacter jejuni. Appl Environ Microbiol 2005;71:987-92. https://doi.org/10.1128/AEM.71.2.987-992.2005
- 6. Lopez AE, Grigoryeva LS, Barajas A, Cianciotto NP. Legionella pneumophila rhizoferrin promotes bacterial biofilm formation and growth within amoebae and macrophages. Infect Immun 2023;91:e0007223. https://doi.org/10.1128/iai.00072-23
- 7. Mengue L, Régnacq M, Aucher W, et al. Legionella pneumophila prevents proliferation of its natural host Acanthamoeba castellanii. Sci Rep 2016;6:36448. https://doi.org/10.1038/srep36448
- 8. Li P, Vassiliadis D, Ong SY, et al. Legionella pneumophila infection rewires the Acanthamoeba castellanii transcriptome, highlighting a class of sirtuin genes. Front Cell Infect Microbiol 2020;10:428. https://doi.org/10.3389/fcimb.2020.00428
- 9. Price C, Jones S, Mihelcic M, Santic M, Abu Kwaik Y. Paradoxical pro-inflammatory responses by human macrophages to an amoebae host-adapted Legionella effector. Cell Host Microbe 2020;27:571-84.e7. https://doi.org/10.1016/j.chom.2020.03.003
- 10. Kim MJ, Jo HJ, Quan FS, et al. Identification of essential genes for Acanthamoeba castellanii excystation during encystation and excystation. Parasites Hosts Dis 2024;62:399-407. https://doi.org/10.3347/PHD.24062
- 11. Moon EK, Chung DI, Hong YC, Kong HH. Autophagy protein 8 mediating autophagosome in encysting Acanthamoeba. Mol Biochem Parasitol 2009;168:43-8. https://doi.org/10.1016/j.molbiopara.2009.06.005
- 12. Song SM, Han BI, Moon EK, et al. Autophagy protein 16-mediated autophagy is required for the encystation of Acanthamoeba castellanii. Mol Biochem Parasitol 2012;183:158-65. https://doi.org/10.1016/j.molbiopara.2012.02.013
- 13. Moon EK, Hong Y, Chung DI, Kong HH. Cysteine protease involving in autophagosomal degradation of mitochondria during encystation of Acanthamoeba. Mol Biochem Parasitol 2012;185:121-6. https://doi.org/10.1016/j.molbiopara.2012.07.008
- 14. Moon EK, Chung DI, Hong YC, Kong HH. Characterization of a serine proteinase mediating encystation of Acanthamoeba. Eukaryot Cell 2008;7:1513-7. https://doi.org/10.1128/EC.00068-08
- 15. Moon EK, Kong HH. Short-cut pathway to synthesize cellulose of encysting Acanthamoeba. Korean J Parasitol 2012;50:361-4. https://doi.org/10.3347/kjp.2012.50.4.361
- 16. Moon EK, Kim MJ, Lee HA, Quan FS, Kong HH. Comparative analysis of differentially expressed genes in Acanthamoeba after ingestion of Legionella pneumophila and Escherichia coli. Exp Parasitol 2022;232:108188. https://doi.org/10.1016/j.exppara.2021.108188
- 17. Aung JM, Joo SY, Na BK, et al. Establishing a Cre/loxP-based genetic manipulation system for Acanthamoeba: targeted genome editing and stable reporter expression. Parasites Hosts Dis 2025;63:25-36. https://doi.org/10.3347/PHD.24078
- 18. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001;25:402-8. https://doi.org/10.1006/meth.2001.1262
- 19. Marti M, Hehl AB. Encystation-specific vesicles in Giardia: a primordial Golgi or just another secretory compartment? Trends Parasitol 2003;19:440-6. https://doi.org/10.1016/s1471-4922(03)00201-0
- 20. Herman E, Siegesmund MA, Bottery MJ, et al. Membrane trafficking modulation during Entamoeba encystation. Sci Rep 2017;7:12854. https://doi.org/10.1038/s41598-017-12875-6
- 21. Rolando M, Escoll P, Buchrieser C. Legionella pneumophila restrains autophagy by modulating the host's sphingolipid metabolism. Autophagy 2016;12:1053-4. https://doi.org/10.1080/15548627.2016.1166325